Note: Descriptions are shown in the official language in which they were submitted.
1325~7~
.
~ Method of Monitoring Cell Culture
,. . _ . _ _ . . .
The present invention relates to a method of
monitoring cell cultures. More particularly, it relates
to a method of monitoring cell cultures which comprises
digi~izing a cultured cell image inputted to image
processing equipment and measuring the number of living
cells, the number of dead cells and the cell proliferation
by processing with a spatial filter.
When cells are cultivated in a vessel, e.g. a
microplate, dish or culturing bottle, some of the living
cells die if the concentration of the living cells become
so high as to reach the so-called "full growth" state. To
prevent this, it is necessary to change the culture medium
and/or to subculture the cells before the full growth state
is reached. ~itherto, a technician determines the timing
for changing the cell culture solution or for starting the
subculturing by observing the cell proliferation of the
j cells in the vessel through a microscope. However, such a
conventional method is not quantitative and th~refore not
accurate in determining the cell proliferation of cultured
cells and the concentration of living cells. In addition,
- lt is time-consuming and it is difficult to precisely
observe the cell proliferation of a large amount of
cultured cells.
~, " ,
,: . . . ~
:. ~ ~ , , . , :
.
.
- 2 - ~32~7~
Further, when cells are cloned by a limiting
dilution-culture method or when cells are fractionated,
both the number of living cells and the number of dead
cells should be simultaneously measured. In such cases,
even if a blood cell counter is used, visual measurement
with the microscope includes large errors and takes a lot
of time.
A cell detection method using image processing has
been proposed instead of visual observation with a micro-
scope. However, it has low detection efficiency as it is
done simply by binary digitizing. Although detection of
the cells is easy when the background is simple, for
example, in the case of an agar medium on which colonies
are formed, when the pattern of cells and the background
are both comlicated, for example, when the cells in the
culturing vessel contain dead cells, dust or other foreign
substances, detection is diEficult.
An object of the present invention is to provide a
method of monitoring cell cultures so that the number of
cells can be precisely measured when the pattern of the
celIs and the background are complicated.
Another object of the present 1nvention is to
provide a method of monitoring cell cultures which can
treat a large number of cells in a short time.
, .
,.
-, : ,
,~
,
_ 3 _ 132~75
In one preferred embodiment of the present invention
these and other objects are achieved by a method of monitoring
: cell culture comprising the steps of: digitizing an image of
. cultured cells which have been input to image processing
- 5 equipment through a TV camera connected with observing means;
extracting only living cells by processing with a spatial
filter having a round shape, a designated size and positive
coefficients in circumferential area wherein said round shape
and designated size is determined from an average pattern of
cell brightness distribution patterns, and measuring the
number of living cells, wherein the cell culture comprises
attachment-dependent eukaryotic cells.
In another embodiment of the invention there is
provided a method of monitoring cell culture comprising the
steps of: digitizing an image of cultured cells which have
been input to image processing equipment through a TV camera
connected with observing means; extracting living cells and
dead cells by processing with a spatial filter having a round
shape, a designated size and positive coefficients in
circumferential area wherein said round shape and designated
size is determined from an average pattern of cell bri~htness
distributed patterns, and measuring the number of living cells
and the number of dead cells, wherein the cell culture
comprises attachment-dependent eukaryotic cells.
In the drawings which illustrate preferred
embodiments of the present invention,
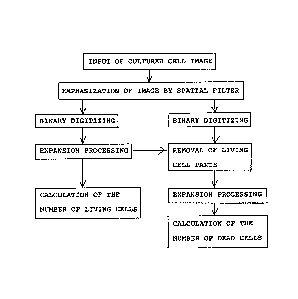
Fig. 1 is a flow chart of procedures for measuring
the number of living cells;
: ~ . . '.,
"
'' ' , " ~. ' " '
~32~7~
; - 3a -
Fig. 2 is a flow chart of procedures for measuring
the number of living cells and the number of dead cells;
Fig. 3 shows shape and coefficients of a spatial
filter having 7 x 7 pixels;
Fig. 4 shows shape and coefficients of an optimized
spatial filter having 7 x 7 pixels; and
Figs. 5 and 6 respectively show shape and
coefficients of spatial filters used in Examples 1 and 2.
-I The method of monitoring cell cultures according to
the present invention will be illustrated by reference to the
accompanying drawings.
By using a TV camera connected to a microscope as an
observing means, an image of the cultured cells in a culturing
vessel is input to image processing equipment. According to
the procedures shown in Fig. l, the image is processed and the
, number of living cells counted.
;1
~,,
~,
I
I
;,
',:
:,
' , ,.:
..... . .
_ 4 _ ~ 32~7~
Preferably, expansion processing is carried out after the
binary digitizing. According to the procedure as shown in
Fig. 2, the number of living cells and the number of dead
cells are measured.
By displacing the focus from an entirely focused
state, the image of the cultured cells is inputted at a focus
point where the center area of the cultured living cells is
brighter and the circumferential area thereof is darker so
that the image has a clear profile. By displacing the focus
of the microscope at a distance of ~5 to -~250 micrometers,
the image of the living cell has a clear profile. When the
displaced distance is less than 5 micrometers, the whole
image of the living cell remains dark. When the displaced
distance is more than 250 micrometers, the image is blurred.
In this state, when the image of one cultured liv-
ing cell is magnified by such magnification that the cellimage inscribes a square of a spatial filter matrix having a
designated size, that is, when the spatial filter matrix has
the size of n x n pixels (n 25), the image of the cultured
cell is inputted at such magnification that one pixel has a
size of 10/n x 10/n to 21/n x 21/n (micrometer x micrometer)
since one cultured living cell generally has a size of 10 to
21 micrometers. The detection efficiency is poor when the
living cell is larger or smaller than the spatial filter
matrix of a designated size. A spatial filter matrix having
a size not smaller than 5 x 5 pixels is used.
,
.. .
" : '' '
,; ' ~,
: '
, , : ,: . .. . `
',
- s- ~325~75
When a spatial filter matrix having a size smaller than S
x 5 pixels is used, the filter has poor detection
efficiency and cannot be used practically.
When the whole image is filtered through a round
shaped spatial filter having negative coefficients in its
circumferential area, positive coefficients in its center
area and zero coefficients in other intermediate areas, the
living cell parts can be emphasized. For example, the shape
and coefficients of the spatial filter having 7 x 7 pixels
are shown in Fig. 3.
The coefficients and shape of the filter for em-
phasizing the living cell parts can be optimized by collect-
ing brightness distribution patterns of the images of plural
objective living cells and determining an average pattern~
The reason for the above is that since the spatial
filtering emphasizes the parts which have better correlation
with the shape of the spatial filter (standard pattern), the
best correlation is realized when the shape of the spatial
filter coincides with the cell brightness pattern. For
example, the shape and coefficients of the optimized spatial
filter having 7 x 7 pixels are shown in Fig. 4.
In this case, a value Yij of the center pixels is
calculated according to the following equation:
1 ~3 +3
Yij A ~ ~ hmn Xi+m,j+n
m=-3 n=-3
wherein x is a value of the pixel before filtering,
, ,
~: : ,
- 6 - 132547~
y is a value of the pixel after filtering,
; A is a coefficient, and
hmn is a coefficient corresponding to each pixel
of the filter.
The pixels are binary digitized with a suitable threshold.
The pixel ir. which the living cell is presen~ is digitized
to be "1" and the pixel in which no living cell is present
is digitized to be "O". The optimized binary digitized
threshold is determined so that in one image, a visually
counted number or the actual number of the living cel.ls
coincides with the number of masses of the pixels which
are digitized as "1" and considered to have the living
. cell, and an average value of those in plural images can
. be used as a fixed binary digitized threshold. The reason
for this is that the correlation of the shape is observed
~; irrespective of background brightness. The number of the
~: living cells is calculated from the number of pixels
i considered to have the living cell (the number of the
;~; pixels which are digitlzed as "1"). Since the number of
pixels which one living cell occupies is constant, the
number of the living cells is calculated according to the
following equation:
The number of living cells =
The number of pixels which are
~: considered to have the living cells
, The~ number of pixels which~~o-ne~~~-~
living pixel occupies
' '~
. . .
;~ , . : . .
.'` . :, '
- 7 - ~32~75
It is possible to decrease the errors resulting
when the number of pixels is converted to the number of the
cells, by subjecting the binary digitized image to a
4-neighbour expansion processing, 8-neighbour expansion
processing or combination thereof one or more times before
calculating the number of pixels from the binary digitized
image. By the expansion processing, the images which are
damaged by binary digitizing can be restored, and the
number of pixels which one cell occupies is increased so
that the influence of one pixel is decreased.
The image emphasized by spatial filtering is
binary digitized again with a threshold different from,
usually lower than, the threshold which is used to deter-
mine the number of living cells and then inverted. Tbe
pixel in which the dead cell is present is digitized to be
"1" and the pixel in which no dead cell is present is
digitized to be 1~0l~. From the number of pixels which are
considered to have the dead cell (the number of pixels
which are digitized as "1"), the number of dead cells is
calculated. Since the num~er of pixels which one dead
cell occupies is constant, the number of dead cells is
calculated according to the following equation:
The number of dead cells=
The number of pixels which are
considered to have the dead cells
,
The num~er of plxels whlcn one
dead cell occupies
.~ , , ; ~ ,. .. .
. , . , :
.
- 8 - ~32~75
The circumferential area (profile~ of the living
cell may also be contained in the pixel of the dead cell
depending on the absolute value of the threshold. This is
removed by a procedure wherein the spatially filtered image
is binary digitized with the threshold for determining the
number of dead cells and then inverted r and a binary digi-
tized image which is binary digitized with the threshold
for determining the number of living cells or a binary
digitized image which is subjected to an expansion pro-
cessing for an appropriate number of times is deducted from
said inverted image on the display. The expansion pro-
cessing is preferably carried out several times. When the
expansion processing is carried out too often, the neces-
sary image is erased. In the same manner as for the living
cells, the error caused when the number of cells is calcu-
lated from the number of pixels is reduced by subjecting
the binary digitized image from which the noise of the
circumferential area of the living cell is removed to a
4-neighbour expansion processing, 8-neighbour expansion
processing or combination thereof one or more times.
Now, the present invention is explained by the
following examples.
;,
.~ .
.. ~
)
.
~ ::., , -, "
., , . ~ , ~ :: . ,
:,. : ~ ' ' : ' I
- 9 - 1~ 2 ~ 47 ~
Example 1
A complicated image in which colonies of cultured
living cells were present together with dead cells, foreign
substances and the like was inputted to image processing
equipment PIAS-l* (available from PC Systems, Japan~
through a microscope connected to a CCD camera by dis-
placing the focus a distance of 30 micrometers from the
entirely focused state.
The magnification was such that the profile of
cultured living cell inscribed a spatial filter having
7 x 7 pixels, that is, one pixel had a size of 1.8
micrometers x 1.8 micrometers.
When the image was digitized with the image pro-
i cessing equipment PIAS-l and filtered with a spatial filter
having shape and coefficients shown in Fig. 5, detection
rate was 67 % and 70 % with an erroneous detection rate of
10 % and 20 %, respectively. The detection rate and the
, erroneous detection rate were determined according to the
following equations:
Detection rate =
The number of actually present living cells
which are detected by the image processing
:
The number of living cells actually present in
the original image
Erroneous detection rate =
` The number of living cells considered to
be present by the image processing but not
actually present in an original image
The number of all living cells which are consi-
dered to be present by the image processing
*Ir~de Mark
'1.
:
," ~ .
~' ' ;. ~ ` '
' ' , .' ' ' ":
. ' ' ' ~ . ' ' ,,, ~.' ' " ``,'' "'' ` '' '
- lo ~2~7~
Example 2
A complicated image in which colonies of cultured
living cells were present with dead cells, foreign sub-
stances and the like was inputted to image processing
equipment PIP-4000* (available from ADS Limited, Japan)
through a microscope connected to a CCD camera by
displacing the focus a distance of 30 micrometers from the
entirely focused state.
` The magnification was such that a profile of
! cultured living cell inscribed a spatial filter having
7 x 7 pixels, that is, one pixel had a size of 2.4
micrometers x 2.4 micrometers.
The image was digitized with the image processing
equipment PIP-4000 and filtered with the spatial filter
having shape and coefficients shown in Fig. 6. When the
~; binary digitizing threshold which is based on the visually
counted number of living cells was used, the detection rate
was 73% and 81% with the erroneous detection rate of 10~
and 20%, respectively. When the same image was subjected
to the binary digitizing alone, the detection rate was 57%
and 60% with the erroneous detection rate of 10% and 20%,
respectively.
! The image in which living cells and dead cells
were randomly present was inputted in the same manner as
above, and the number of living cells and the number of
! dead cells were measured. As for the living cells, the
.
, *Trade mark
)
:, . , . - . , .
..
L3~5475
visually counted number was 47 and the number measured by
the image processing was 50, an error of 6.4%. As for the
dead cells, the visually counted number was 453 and the
number measured by the image processing was 453, an error
of 0~. Further, the error of the measured number of total
cells (the number of living cells and the number of the
dead cells) had a standard deviation of +10.2~ in one sigma
for 39 samples. The error was determined according to the
following equation:
Error =
; (The number measured by image processing) -
(The visually counted number)
The visually counted number
Example 3
The procedure as in Example 2 was repeated except
that the binary digitizing threshold based on the actual
number of living cells was used instead of the binary
digitizing threshold based on the visually counted number of
~' living cells. When the image was filcered with the
spatial filter having the same shape and coefficients as
used in Example 2, the detection rate was 73 % and 82 ~ in
the erroneous detection rate of 10 % and 20 %, respectively.
When the binary digitized image was subjected to
8-neighbor expansion processing once, the error caused by
`~ converting the number of pixels to the number of
, living cells was 8 ~. It was 17 % when the image had not
been subjected to the expansion processing.
~' .
, .. , :. . . :
; . . :,. . .
: . .