Note: Descriptions are shown in the official language in which they were submitted.
WO 96/09381 PCT/FI95/00520
- 1 -
PROMOTER FOR THE RECEPTOR TYROSINE RINASE, TIE
FIEhD OF THE INVENTION
The present invention relates generally to
receptor tyrosine kinases and promoters thereof.
BACKGROUND OF THE INVENTION
The circulatory system is the first organ
system to differentiate in the developing embryo.
Kaufman, The Atlas of Mouse Development, Academic
Press, (1992). Embryonic and yolk sac vascular
systems take form in an 8.5 day p.c. mouse embryo
and a day later the heart beats regularly,
circulating primitive blood cells, nutrients, and
metabolic waste products. Endothelial cells covering
blood vessels provide a barrier between blood and
other tissues of the embryo. When organs
differentiate and begin to perform their specific
functions, the phenotypic heterogeneity of
endothelial cells increases. Fenestrated vessels,
nonfenestrated vessels with tight junctions and
sinusoidal vessels are found, for example, in the
kidney, brain, and liver, respectively. In addition,
endothelial cells perform specific functions in
differentiated tissue. For example, such cells take
part in several biochemical and physiological events
such as blood cell trafficking, blood clotting,
hemostasis, ovulation, wound healing,
atherosclerosis, and angiogenesis associated with
tumor metastasis.
At least five receptor tyrosine kinase
genes are expressed in endothelial cells. Of these,
the protein products of the FLT1, KDR/FLK-1, and
FLT4 genes belong to receptor tyrosine kinase
subclass III; whereas Tie and its close relative Tek
SUBSTITUTE SNEET RULE 26)
WO 96/09381 P~.'TIFI951GOSIa
~IvIN~~
- 2 -
(Tie-2) form a novel subclass of their own (Terman,
et al., Oncogene, 6: 1677-1683, 1991, Terman, et
al., Biochem. Biophys. Res. Comm., 187: 1579-1586,
1992, Aprelikova, et al., Cancer Res., 52: 746-748,
1992, De Vries, et al., Science, 255: 989-991, 1992,
Pajusola, et al., Cancer Res., S2: 5738-5742, 1992,
Sarzani, et al., Biochem. Biophys. Res. Comm., 186:
706-714, 1992, Galland, et al., Oncogene, 8:
1233-1240, 1993, Millauer, et al., Cell, 72:
835-846', 1993, Oelrichs, et al., Oncogene, 8: 11-18,
1993, Schnurch and Risau, Development, 119: 957-968,
1993). Both human and mouse Tie cDNAs have been
cloned (Partanen, et al., Mol. Cel. Biol., 12:
1698-1707,1992, Korhonen, et al., Blood, 80:
2548-2555, 1992, Korhonen, et al., Oncogene, 8:
395-403, 1994, Iwama, et al., Biochem. Biophys. Res.
Comm., 195: 301-309, 1993, Sato, et al., Proc. Natl.
Acad. Sci. USA., 90: 9355-9358, ,1993). Tie and
homologous genes have been isolated from bovine and
rat sources (Maisonpierre, et al., Oncogene, 8:
1631-1637,1993, Sato, et al., Proc. Natl. Acad. Sci.
USA., 90: 9355-9358,:1993). Genomic clones for
mouse Tie and both mouse and human Tie promoter
regions have been cloned and characterized.
The 4.4 kb Tie-encoding mRNA encodes a 125
kDa transmembrane protein which is N-glycosylated.
In its extracellular domain Tie contains two
immunoglobulin-like loops and three epidermal growth
factor and fibronectin type III homology regions,
which are followed by traps- and juxtamembrane
domains connected to a tyrosine kinase domain which
is split by a short kinase insert sequence and a
carboxyl terminal tail (Partanen, et al., Mol. CeI.
Biol., 12: 1698-1707, 1992, Korhonen, et al.,
Oncogene, 8: 395-403, 1994, Sato, et al., Proc.
a
SItgSTITUTE SHEET (RULE 26)
WO 96/09381 ~ ~ ~ 4 ~ PCT/FI95100520
- 3 -
Natl. Acad. Sci. USA., 90: 9355-9358, 1993). Both
Tie and TEK have been localized to mouse chromosome
4 at a distance of 12.2 cm from each other. Such
receptors are uniformly expressed in endothelial
cells of various blood vessels during embryonic
development, although the expression of Tek mRNA
appears to begin 0.5 days earlier than the
expression of Tie. In adult mice, the expression of
Tie mRNA persists in vessels of the lung whereas in
the heart and brain it appears to decrease.
Korhonen, et al., Oncogene, 8: 395-403, (1994).
Production of Tie mRNA is enhanced during ovulation
and wound healing and in human glioblastomas
(Korhonen, et al., Blood, 80: 2548-2555, 1992).
Endothelial cells play a key role in gene
therapy directed to diseases involving endothelial
cells and blood vessels, such as establishment of
neovascularization or inhibition of angiogenesis,
and control of inflammatory trafficking of
leukocytes. One approach to the treatment of
vascular disease is to express genes at specific
sites in the circulation that might ameliorate the
disease in situ. Because endothelial cells are
found at diseased sites, they represent logical
carriers to convey therapeutic agents that might
include anticoagulant, vasodilator, angiogenic or
growth factors. Accordingly, the genetic
modification of endothelial cells represents a
therapeutic approach to the treatment of many
vascular disorders, including hypertension,
atherosclerosis and restenosis. For example,
endothelial cells expressing growth inhibitory
' proteins could be introduced via catheter to the
angioplasty site to prevent local intimal
hyperplasia and clinical restenosis. The luminal
SUBSTITUTE SHEET (RULE 26~
WO 96109381 ~ PC.TIFi95/0U52U . .
- 4 -
surface of vascular grafts could also be lined with
genetically modified endothelial cells producing
therapeutic proteins which prevent thrombosis or
promote repooulation (Nabel, et al., J. Am. Coll.
Cardiol., 17, 189B-194B, 1991).
Endothelial cells lining blood vessels are
easily transfected with methods using liposomes,
adenovirus vectors and retroviral vectors (Nabel, et
al., J. Am. Coll. Cardiol. 17: 189B-94B).
l0 Endothelial cells are also in direct contact with
blood and are therefore optimal sources for
production and secretion of desired proteins or
peptides into the blood stream. For example, the
Factor VIII gene may be introduced into endothelial
cells under an endothelial cell- specific promoter,
resulting in correction of hemophilia if the protein
were expressed in sufficeint quantity. on the other
hand, endothelial cells are also useful for delivery
of peptides or proteins expressed in them into
tissues. In this regard, a selective expression of
a particular gene regulatory element in endothelial
cells of the microvasculature (capil.laries) is
extremely useful; given that most of the cell
surface area facing the vascular lumen consists of
microvascular endothelial cells.
Control elements of the endothelial cell
specific promoters may be further subdivided and
dissected into functional elements and units
according to methods standard in the art. The Tie
3o protein is expressed in certain endothelial cells
and about 0.9% of human bone marrow cells.
Therefore, it is likely that the Tie promoter is
active also in some hematopoietic cells. However,
expression of the Tie promoter in hematopoietic
cells may be controlled by elements which are
f
SUBSTITUTE SNEET (RULE 26~
WO 96/09381 ~ P'~.'T/FI95/005~0'
- 5 -
distinguishable from endothelial- cell-specific
elements and may be dissected away while retaining
the endothelial cell specificity of the promoter.
The present invention provides a novel
promoter associated with the gene encoding the Tie
receptor tyrosine kinase for use in therapeutic and
diagnostic procedures. In addition, the promoter may
prove useful in the production of desired proteins
to the blood or tissues of animals.
SUMMARY OF THE INVENTION
The present invention generally relates to
promoter sequences for the receptor tyrosine kinase,
Tie. In a preferred embodiment of the invention, a
mouse Tie promoter is provided comprising the
sequence shown in SEQ ID NO:1. Also in a preferred
embodiment, a human Tie promoter is provided
comprising the sequence shown in SEQ ID N0:2. A
promoter according to the invention drives the
expression of endothelial cell receptor tyrosine
kinases, and in particular, the receptor tyrosine
kinase, Tie.
~ vector according to the present
invention may be any vector suitable for
incorporating a promoter according to the invention
, and may preferably be the 0.73mTIEpromGL2 vector
deposited on September 19, 1994 with the American
Type Culture Collection, 12301 Parklawn Drive,
Rockville, MD 20852, as Accession Number 75892
Host cells according to the invention :nay be any
host cell capable of housing the promoter or a
vector containing the promoter according to the
invention. Examples of host cells according to the
invention are LEII endothelial cells.
0
SUBSTITUTE SHEET (RULE 26)
WO 96/09381 , ~ ' PC'T/F79510~S2u . ~ , .
- 6 -
Other advantages and uses of the invention
will be apparent upon consideration of the following
Detailed Description thereof.
DESCRIPTION OF THE DRAWINGS
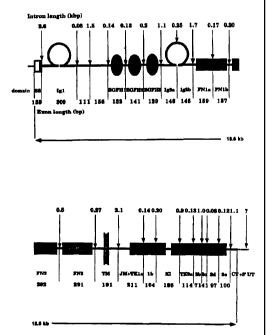
Figure 1 is a schematic diagram of the
mouse Tie gene and promoter.
Figure 2 is a comparison of mouse and human
Tie promoter sequences. (SEQ ID NO:1 and SEQ ID N0:2).
Figure 3 shows results of an analysis of
Tie promoter activity.
Figures 4A shows expression patterns of
the Tie promoter in the developing endocardium and_
head mesenchyme of 8.5 day mouse embryos.
Figure 4B show expression of a mouse Tie
promoter construct in yolk sac blood islands in 8.5
day embryos.
Figure 5A and 5B show the expression
pattern of mouse Tie promoter in 9.5 day embryos.
Figures 5C and 5D show the expression
pattern of mouse Tie promoter in 11.5 day embryos.
Figure 6A shows expression of the Tie
promoter in-9.5 day embryonic heart tissue.
Figure 6B shows expression of the Tie
promoter in 11.5 day embryonic lung tissue.
; Figure 6C shows expression of the Tie
promoter in 15.5 day embryonic brain tissue.
Figure 6D shows expression of the Tie
promoter in 15.5 day embryonic liver tissue.
Figure 6E shows expression of the Tie
promoter in developing bone trabeculae.
Figure 6F shows expression of the Tie
promoter in developing kidney tissue.
SUSST1TUTE SHEET (RULE 26)
WO 96/09381 ~ ~ ~ ~ ~ PGT/FI95I00520
-
Figure 7A shows expression of the Tie
promoter in the interalveolar capillaries of the
lung in an 8-week-old mouse.
Figure 7B shows expression of the Tie
promoter in the endothelial network of the bone
marrow in an 8-week-old mouse.
Figure 7C shows expression of the Tie
promoter in kidney tissue of an 8-week-old mouse.
Figure 7D shows expression of the Tie
promoter in heart tissue of an 8-week-old mouse.
Figure 7E shows expression of the Tie
promoter in liver tissue of an 8-week-old mouse.
Figure 7F shows expression of the Tie
promoter in brain tissue of an 8-week-old mouse.
DETAINED DESCRIPTION OF THE INVENTION
The present invention provides promoter
sequences capable of directing the expression of
recombinant DNA sequences in endothelial cells. In
particular, the invention provides promoter
sequences which direct expression of the
beta-galactosidase reporter gene in endothelial
cells of mouse tissues. Promoters for production of
proteins and peptides which act as anticoagulants,
vasodilator inhibitors of thrombosis or restenosis
into endothelial cells, blood and tissues.
Promoters according to the present invention are
useful for directing expression of proteins and
peptides for human gene therapy, antigens and
markers useful for endothelial cell tagging, and
antisense RNA constructs for use in endothelial
cells in vivo and in vitro. Promoters, vectors, and
host cells according to the invention are also
useful in gene therapy for promoting expression of
various growth factors or receptors or their
SUBSTITUTE SHEET (RULE 26)
~p~~46
WO 96/09381 PCT/F'I95100520
_ g
domains. Moreover, analogs of promoters according to
the invention are useful for inhibiting undesired
endothelial cell proliferation as, for example, the
inhibition of angiogenesis during tumor formation.
EXAMPLE I
Cloning and characterization of the
genomic Tie DNAs.
A. Mouse Genomic Tie
In order to characterize the genomic
organization of the mouse Tie gene, approximately 3
x 106 plaques were screened. The plaques were
obtained from a genomic library made from DNA of
adult SV129 mouse liver cells (Clontech) using as a
probe a mouse 1C1D cDNA fragment (Korhonen, et al.,
Blood., 80:2548-2555, 1992) encoding the epidermal
growth factor homology domains
[GCVKDCPGCLHGGVCHDHDGCVCPPGFTGTRCEQACREGRFGQSCQEQCPG
TAGCRGLTFCLPDPYGCSCGSGWRGSQCQEACAPDHFGADCRLQCQCQNGGT
CDRFSGCVCPSGWHGVHCEKSDRIPQIL: SEQ ID N0:3] Three
separate clones, SV1, SV2, and mTie were obtained
thereby and each was subcloned into pGEM 3Zf(+)
(Promega) and characterized by partial dideoxy chain
termination sequencing and restriction enzyme
analysis. A schematic structure of the mouse Tie
gene and its promoter is shown in Figure 1. In that
Figure, the positions of introns are indicated by
arrows and their lengths are indicated. Restriction
mapping, PCR, and nucleotide sequence analysis
showed that the Tie gene spans approximately 19 kb
of genomic DNA. Tie is encoded by 23 exons. The
distinct structural domains of the,extracellular
portion are encoded by either one exon each,
comprising the first immunoglobulin-like loop,
epidermal growth factor homology domains 1-3 and
SUBSTITUTE SHEET (RULE 26j
CA 02200846 2005-05-20
zzaas~s : :..
. . .
WO 96/09381 . ~ Pt,'T1FI9516051b.
- g _
f ibronectin-like domains 2 and 3, or by two exons
comprising the second immunoglobulin-like loop and
first fibronectin-like domain. The transmembrane
region is encoded by a distinct exon; whereas the
tyrosine kinase domain containing the kinase insert
is encoded by eight exons of which the first encodes
the juxtam~mbrane region. The lengths of the
introns vary from 80 by to 2.6 kb.
H. Human Genomic Tie
Three human Tie clones were isolated from
a human placental genomic DNA library in the EI~L-3 vector
system (Clontech) as shown in Partanen, et al., Mol.
Ce~l. Eiol., 12: 1698-1707 (1992),
To obtain the human Tie clones, a
PCR fragment encoding the Tie signal sequence was
amplified from human Tie cDNA using the primers,
5'-CCCACATGAGAAGCC-3' (SEQ ID NO:. 4) and
5'-TGAGATCTTGGaGTATGGTCTGGCGGGTGCCC-3' (SEQ ID NQ:
5), and used to probe the aforementioned library.
The resulting positive clone containing the longest
insert was plaque-purified and an approximately 7 kb
SacI fragment was subcloned in pGEM 3Zf(+) and
characterized. The resulting human Tie promoter
sequence is shown in Figure 2. In that Figure,
? transcription initiation sites are marked with an
asterisk (See primer extension and RNAse protection
experiments below). Restriction endonuclease
cleavage sites discussed herein are marked in bold.
A comparison of the genomic DNA sequences
of mouse and human Tie promoters is also shown in
Figure z. In that Figure, the mouse sequence
extends from the 3' end of the first~exan to the
AfZII site which is approximately 821 by upstream
a
SUBSTITUTE SHEEP (RULE 26)
.~ .-'- ,'- y
WO 96/09381 , ' ,PCT/FI95/~52~
- 10 -
from the ATG cvdon. A CA repeat found only in the
mouse sequence is highlighted in bold in Figure 2.
EXAMPLE II
Determination of The Transcription Initiation Site
in The Human Tie Gene
For primer extension analysis of
Tie-encoding nucleic acids, primer was labelled
according to the manufacturer's instruction
(Promega, USA). An aliquot of l0 pmol primer was
then incubated with 10 x forward exchange buffer
(Promega) , 10 ~,Ci/ml ['y-3zP] -ATP, and l0U T4
polynucleotide kinase at 37°C for 1 hour. The
kinase was then inactivated by heating at 90°C for 2
minutes and the labelled primer was ethanol
precipitated.
Poly (A+) RNA (20 ~,g) and 5 x 105 cpm
labelled primer were then annealed in hybridization
buffer (40 mM PIPES pH 6.4, 1mM EDTA pH 8.0, 0.4 M
NaCl and 80% formamide) by heating at 95°C for 12
minutes. Samples were then cooled slowly and ethanol
precipitated. The resulting dried annealing mixture
was suspended in primer extension buffer (SO mM
Tris- HC1, 50 mM KC1, 10 mM MgCl2, 10 mM DTT, 2 mM
each of deoxy ATP, deoxy CTP, deoxy GTP, and deoxy
~ TTP, 0.5 mM spermidine, pH 8.3, at 42°C) and 20 U
RNAsin and 40 U AMV reverse transcriptase were
added. After 2 hours of incubation, template RNA was
digested by addition of 20 ~g/ml RNAse A in 100 mM
NaCl, 10 mM Tris-HC1, 1 mM EDTA, pH 7.4, 37°C for 15
minutes. The resulting mixture was phenol extracted
and ethanol precipitated. The pellet was then
resuspended in loading dye (980 formamide, 10 mM
EDTA, O.lo xylene cyanol, 1% bromophenol blue) and
loaded onto a 9% polyacrylamide/7M urea gel. After
p f
SUBSTITUTE SHEET (RULE 26)
~s - ' " ,
WO 96/09381 . ~ PGTIFT95/Q952~
., ...
- 11 -
electrophoresis, the dried gels were exposed to
x-ray film for 2 days.
RNAase protection was accomplished using
mouse RNA antisense probes of 291 by and 239 by gene-
s rated from linearized plasmids containing the approx-
imately 842 by AflII - BamHI and 1.1 kb HindIII - ApaI mouse
Tie promoter DNA inserts. The human RNA probe of 568
by was generated from linearized pGEM 3Zf(+) plasmid
(Promega, USA) containing an AccI-AlwNI human Tie
l0 promoter DNA insert. The template for the other
human 266 by RNA probe was generated by PCR
amplification from the AccI-AlwNI plasmid. M13
Forward and Tie 2168 primers (marked in Figure 2)
were used for amplification. The probes were labeled
15 using T7 polymerase and ('y-32PJ -UTP. 10 ~g of poly
A(+) RNA was incubated with labelled probe at 50°C
overnight. Unhybridized RNA was digested with RNAse
A (10 U/ml) and T1 (1 ~Cg/ml) at 37°C, pH 7.5 for 1
hour. The RNAses were inactivated by proteinase K
20 digestion at 37°C for 15 minutes and the samples
were analyzed in 8% sequencing gels.
The primer~extension and RNase protection
products terminated at positions 101 by and 116 by
upstream from ATG codon, in mouse and human Tie
25 promoters, respectively (see asterisks in Figure 2).
? Yeast tRNA or NIH 3T3 RNA did not show any specific
bands. Results are shown in Figure 2, wherein the
sequences of primers referred to above are
underlined.
30 EXAMPhE III
Construction of plasmids
Tie promoter/luciferase gene constructs
were generated by subcloning the 5' flanking approxi-
SU8ST1TUT~ SHEET (RULE 26)
CA 02200846 2005-05-20
.~ .
WO 96/U9381 P~,~~~5~ . ' . _ . ;
... . .. '
- 12 -
mat a ly 788 by genomic AflII-ApaI fragment located upstream
of the ApaI restriction site in the first exon to the
promoterless basic pGL2 vector (Stratagene) as
described by deWet, et al., Mol. Cell. Biol., 7:
725-737 (1987),
resulting in plasmid 0.73mTIEpromGL2.
For experiments in transgenic mice, the ap-
proximately 788 by AfIII-ApaI promoter fragment (shown in
Figure 2) was blunt-end ligated into a blunted, unique
HindIII site in the SDK-LacZ Hluescript vector
(Stratagene), as described in Logan, et al.,
Development, 117: 905-916 (1993)
resulting in vector 0.73mpromSDK-
LacZ. Similarly the 5 kb AlwNI fragment of the
human Tie promoter shown in Figure 2 was blunt-end
ligated into that same vector, resulting in plasmid
S.OhTIEpromSDK-LacZ and deposited with the American
Type Culture Collection, 12301 Parklawn Drive,
Rockville, MD 20852, as Accession Number 75893
~XAMPL~ IV
DNA transfection and preparation of cell lysates
15 ug of the 0,73mTIEpromGL2 plasmid
described above was transfected into either LE II
25? mouse lung endothelial cells which are described in
Schrieber, et al., Proc. Natl. Acad. Sci. (USA), 82:
6138-6142 (1985)
or 2~C-2 cells described by weissman, et al., Cell,
32: 599-606 (1983),
Transfection was accomplished using the
modified calcium phosphate mediated transfection
method reported in Sambrook, et a1. (eds.),
Molecular Cloning: A Laboratory Manual (1989),
The DNAs were
4
SUB~STiTUTE SHEET (RULE 26)
CA 02200846 2005-05-20
WO 96109381 ~ ~ ~ ~ ~ ~ ~ ~PCTJ~T95/08520
, . .,. .. ,. ,.
- 13 -
mixed with 0.25 M CaClZ and an equal volume of 50 mM
N,N-bis(2-hydroxyethyl)-2-aminoethane sulfonic
acid-buffered saline. The mixture was incubated for
15 minutes at room temperature and then added
dropwise on growing cell monolayers.
The resulting cultures were incubated for
12 hours, after which 5% glycerol in PBS (phosphate
buffered saline) was added for 30 seconds and washed
off with two changes of PHS. Fresh medium was~then
l0 added. After further incubation for 24 hours, the
cells were lysed in 0.5 ml lysis buffer (25 mM
Tris-P04, 2 mM dithiothreitol (DTT), 2 mM
1,2-diamino-cyclohexane, N,N,N',N'-tetracetic acid;
l0% glycerol, 1% Triton X-100, pH 7.8). The
resulting lysates were centrifuged and the
supernatants were collected and stored at 70°C until
further assayed. Normalization of luciferase values
relative to transfection efficiency was achieved by
cotransfection of a CMV-,Q-gal vector described in
MacGregor and Casket', Nucl. Acids. Res., 17: 2365
(1989),
Assays for ~-galactosidase and luciferase
were conducted on transfected cells. For the
~i-galactosidase assay, 30 ml of the cell lysate
described above was incubated in 33 ml. of
?o-nitraphenyl-~-D-galactopyranoside (4 mg/ml)
dissolved in 100 ml 0.1 M sodium phosphate, pH 7.5
for 30 minutes at 37°C. Optical density was
measured at 414 nm.
Luciferase assays were performed using a
FlyLight monitoring Kit (102-100, BioTools, Finland)
according to the manufacturer's protocol. Briefly,
20 ml of cell lysate was incubated in 100 ml
reaction mixture and a Bio-Orbit 1253 luminometer
was used to determine light intensity.
SUBSTITUTE SHEET (RULE 26)
_.
WO 96/09381 PCT~SIOe52~ ; ' ' ' . ; o
." , "~ ~,
- 14 -
The activity of the Tie promoter in
cultured cells was measured using Tie
promoter-luciferase constructs described above. The
Tie promoter-luciferase constructs were transfected
into either LEII endothelial cells or into MK-2
epithelial cells. Promoter activity was determined
as the ratio of luciferase to ~i-galactosidase
activity. Those activities were compared to the
promoter activity of the positive control vector,
IO RSV-luc (ATCC).
Activity of the 0,73mTIEpromGL2 plasmid (788)
relative to CMV-,Q-gal, used as a constitutively
expressed cotransfected control promoter, is shown
in Figure 3, along with values for the highly
expressed RSV-luc promoter. A 460 by mouse Tie
promoter fragment was used in reverse orientation as
a negative control (reverse). As shown in figure 3,
the Tie promoter was highly active in LEII cells but
not in the epithelial cells, MK-2. Those results
indicate that the isolated Tie promoter is specific
for vascular endothelial cells and efficiently
promotes the expression of the reporter in those
cells in comparison to the control.
EXAMPLE 0
Production of transgenic mice
The Tie-containing transgene was separated
from the vector sequence by digestion with SalI,
purified by electrophoresis through an agarose gel,
and recovered by absorption on glass beads (Gene
Clean II, Bio 101 Inc., La Jolla, CA) according to
the manufacturer's instructions. Transgenic mice
were produced by the standard microinjection
technique reported in Hogan et al., Manipulating the
a
SU8ST1TUTE SHEET (RULE 26)
CA 02200846 2005-05-20
~zaa~~s
- . . . ; :.
wo 9sro9s8i p~-r~srooszfl ' ' ' ' '
..
... ,
- 15 -
mouse embryo (Cold Spring Harbor, 1986),
Zygotes for
microinjections were obtained from superovulated
(BALB/c X DBA/2)F1 hybrid female mice (CD2F1) mated
with CD2F1 males. Alternatively, eggs used for
injection were from randomly bred superovulated CD1
females. After microinjection, zygotes were
transferred at the one or two cell stage into
oviducts of pseudopregnant foster mothers (CD2F1
to mice). Tail samples were taken from mouse pups at
three weeks of age and DNA Was isolated from the
samples by the salt precipitation method of Miller,
et al., Nucl. Acids. Res., 16: 1215 (1988),
The polymerase
chain reaction was used to confirm.the presence of the
transgene using the mouse promotez-specific primer,
5'-CTATTGAGAAGGTTTGGAGG3-3'[SEQ ID N0:6), the lacZ
vector primer, 5'-GCTCTAGAACTAGTGGATC-3'[SEQ ID
N0:7]; the human promoter-specific primer,
5'-GAGACAGGGGATGGGAAAAA-3' [SEQ ~ID NO:B]; and the
lacZ vector primer, 5'-GAAGATCGCACTCCAGCCAG-3' [SEQ
ID NO: 9] using a reaction mixture comprising 200 ng
DNA (Tail)..; lOx buffer (2mM MgCl2), 250 nM Primer
2040, 250 nM Primer 1986, 0.2 mM dNTP Mixture, 0.02
U Dynazyme(Finnzymes, Finland), and 50 ml of
distilled water, plus 50 ml mineral oil (M-3516;
Sigma, USA). The PCR Program consisted of a hot
start at 96°C, 2 minutes, with cycling as follows:
96°C 1 minutes, 50°C 2 minutes, 72°C 3 minutes, for
34 Cycles, with the last step delayed 10 minutes.
EXAMPLE VI
Analysis of Tie-containing Tissue
Whole mouse embryos were obtained and
stained for (3-galactosidase activity. Tissue was
t
SU8S11TUTE SHEET (RULE 26)
220086
WO 96/09381 PCT/FI95/00520
- 16 -
transferred into 4x paraformaldehyde in PBS (pH 7.4)
and incubated at 4°C for 20 minutes with gentle
agitation. Tissue was then washed with PBS and
incubated in fresh X-Gal reaction mixture [1 mg/ml
4-chloro-5-bromo-3-indolyl-~i-galactoside, 4mM
K4Fe(CN) 6 x 3H20, 2mM MgCl2 in PBS] at 30°C for 1 to
2 days. Then, samples were washed in PBS for 5
hours and transferred to 30% sucrose for storage.
Samples were then embedded in Tissue Tek
(Miles, USA) and 15 ~Cm sections were cut on
silane-treated slides. Sections were post-fixed in
4% paraformaldehyde for 5 minutes, and washed twice
in PBS and once in distilled water. Nuclear fast
red was applied as a counterstain.
Results are provided in Figures 5~and 6.
Figures 5A-5D show expression of the mouse Tie
promoter in 9.5 (Figures 5A and 5B) and 11.5
(Figures 5C and 5D) day post coitum mouse embryos.
As seen in the figures, activity of the
~i-galactosidase reporter gene is found in the
developing heart (h), branchial vessels (ba), paired
dorsal aorta (da), vitelline artery (v), umbilical
artery (u), and in capillaries (c) of 9.5 day post
coitum embryos. Two days later (figures 5C and 5D),
a similar pattern is found with the addition of
staining in the mesonephros (m) and the veins of the
liver(1).
Figures 6A through 6F show Tie promoter
activity in 9.5, 11.5, and 13.5 day post coitum
embryos. All endothelial cells of the cardiac
region are stained, indicating expression under
control of the promoter. Staining is observed in
lung, but the bronchi are negative. Brain tissue of
15.5 day embryos also shows staining. Figure 6D
shows that the promoter is expressed in veins of the
SU85TITUTE SHEET (RULE 26~
PCT/FI95/00520
WO 96/09381
- 17 -
liver and Figure 6E shows staining in the developing
bone trabeculae. The developing cortex of the kidney
shows staining, expression being most prominent in
the glomeruli.
To study the promoter activity during
development, the 0.73mpromSDK-LacZ and
S.OhpromSDK-LacZ DNAs were injected into fertilized
mouse oocytes. Six transgenic mice were obtained,
transgenic males were mated with wild-type NMRI
females and the offspring (86 for 735 by fragment
and 57 for 5.0 kb fragment) were analyzed on days
7.5 - 17.5 of development. Of the F1 offspring, 40%
were positive in LacZ staining, although the embryos
showed a variation of the intensity of the reaction
color. No staining was seen in 7.5 day post-coital
embryos, whereas in 8.5 day post coitum embryos,
endothelial cells of the dorsal aorta and forming
heart were strongly positive. Certain cells of the
head mesenchyme, presumably differentiating
angioblasts, showed a faint signal, and the
extraembryonic tissues, such as allantois and yolk
sac, contained positive vessels.
The complexity of the vascular system
increases rapidly in the developing embryo, and in
9.5 day post coitum embryos promoter activity was
seen in the above mentioned vessels as well as in
the intersomitic arteries. An especially intense
staining was seen in the developing ventricles of
the heart. In 11.5 day post coital embryos the
capillary system is well-developed and therefore the
staining associated with large vessels of the embryo
and the endocardium was only faintly discerned
through the dense network of blue-stained
capillaries. The details of vascular system were
better visualized in high magnification of tissues
SUBSTITUTE SHEET (RULE 26)
;., ~,
WO 96/09381 ~ , , ~ PCT/FT95/ON52(~ ~ ' ' ; ;
.., .. ,.. .,
- 18 -
of day 11.5 and 15.5 post coital embryos. That
staining pattern corresponds to the expression
pattern obtained in in situ hybridization. As
shown in Figures 4A and 4B, the endocardium of the
heart, the veins and the arteries of the head
mesenchyme showed LacZ signal. No significant
differences were seen in the staining patterns
obtained with mouse approximately 788 by and human
5.0 kb promoter fragments.
l0 In order to determine if the promoter ac-
tivity of the approximately ~s8 by mouse fragment cor-
relates with expression of Tie mRrrA in adult tissues,
various tissue types obtained from 8-week old
transgenic mice were stained for ,Q-galactosidase
activity. As shown in Figures 7A and 7B, intense
staining was observed in lung (figure 7A) and bone
marrow (designated bm in Figure 7B). Figure 7B also
shows staining in capillaries associated with hair
follicles (designated by the arrow in Figure 7B).
Slightly less staining was observed in kidney
glomeruli (Figure 7C, designated "g") and vessels
surrounding the tubuli (Figure 7C, designated by the
arrowhead).- Figure 7D shows staining in the
endocardium. Neither large hepatic vessels (v in
Figure 7E) or sinusoidal capillaries (not shown)
stained with LacZ in adult mice. However, small
vessels surrounding the veins did stain [Arrows in
Figure 7(E)]. ~As shown in Figure 7(F), interstitial
capillaries of the brain were stained [Arrowheads
in Figure 7(F)]. Similar results were obtained when
transgenic mice expressed the 5 kb human Tie
promoter.
The present invention has been described
in. terms of its preferred, embodiments. Accordingly,
the invention should be limited only by the scope of
the appended claims.
0
SU8ST1TUTE SHEET (RULE 26)
WO 96/09381 PCT/FI95/00520
19
INDICATIONS RELATING TO A DEPOSITED MICROORGANISM
(PCT Rule l3bis)
A. T'be indicationc~ade below relate
to the microorganism referred
t~~ the description
Gt5
J
, line
.
on page
B. IDENTIFICATION OF DEPOSIT Further
deposits are identified on an
additional sheet
Name of depositary institution
American Type Culture Collection
(ATCC)
Address of depositary institution
(including postel code and country)
12301 Parklawn Drive, Rockville,
MD 20852, USA
Daze of deposit 20 September 1994 ~sion Number ATCC 75892
C. ADDITIONAL INDICATIONS (leave
blank ijnot applicable) This information
is continued on an additional
sheet
In respect of those designations
in which a European patent or
a patent in Finland or Norway
is
sought, a sample of the deposited
microorganism will be made available
until the publication of the
mention of the grant of the European
patent or the corresponding information
concerning the patent in
Finland or Norway or until the
date on which the application
has been refused or withdrawn
or is
deemed to be withdrawn, only by
the issue of such a sample to
an expert nominated by the person
re-
questing the sample (Rule 28(4)
EPC and the corresponding regulations
in Finland and Norway).
D. DESIGNATED STATES FOR WINCH
INDICATIONS ARE MADE (ijtlu indications
are not jor all daignatcd Starts)
E. SEPARATE FURNISHING OF INDICATIONS
(leave blank ijnot applicable)
The indications listed below will
be submitted to the International
Bureau later (specijytJugenerol
natareojtheindications eg., ACf:GiftOn
Nwrrbn ojDeposit
For receiving Office use only For International Bureau use only
~ This sheet was received with the international application ~ This sheet was
received by the International Bureau on:
Authorized ofFtcer _ Authorized officer
~,IGuG: /!~ ~e~ ~t.~..
Form PCT/RO1134 (July 1992)
WO 96/09381 PCT/FI95100520
INDICATIONS RELATING TO A DEPOSITED MICROORGANISM
(PCT Rule 136is)
A. The indicationc~ade below relate
to the microorganism referred
t 1~ the description
J
1G
e
, tin
on page
B. IDENTIFICATION OF DEPOSIT Further
deposiu are identified on an additional
sheet
Name of depository institution
American Type Culture Collection
(ATCC)
Address of depository institution
(including postal code and country)
12301 Parklawn Drive, Rockville,
MD 20852, USA
Date of deposit Accession Number
20 September 1994 ATCC 75893
C. ADDTTIONAL IT1DICATIONS (leave
blank if not applicable) 'Ibis
information is continued on an
additional sbeet
In respect of those designations
in which a European patent or
a patent in Finland or Norway
is
sought, a sample of the deposited
microorganism will be made available
until the publication of the
mention of the grant of the European
patent or the corresponding information
concerning the patent in
Finland or Norway or until the
date on which the application
has been refused or withdrawn
or is
deemed to be withdrawn, only by
the issue of such a sample to
an expert nominated by the person
re-
questing the sample (Rule 28(4)
EPC and the corresponding regulations
in Finland and Norway).
D. DESIGNATED STATES FOR V1~CH
INDICATIONS ARE MADE (i f the
indications are not for oll designated
Stator)
E. SEPARATE FURNISHING OF INDICATIONS
(leave blank if not applicable)
The indications listed belowwill
be submitted to the International
Bureau later (spxifythegaraal
notrtreoftheindicotions cg., 'ACCCSfJ0I1
Number of Deposit
For receiving Office use only For International Bureau use only
~ This sbeet was received with the international application Q This sheet was
received by the International Bureau on:
Authorized offiee~~~ ~ /~~ ~ ~, ~ ~ Authorized officer
(~~t/~~-
Form PCT/RO/134 (July 1992)
WO 96/09381 PCTIFI95/00520
21
Indications relating to deposited microorganisms
Continuation to C. ADDITIONAL INDICATIONS
ATCC 75892 and ATCC 75893
When designating Australia, in accordance with regulation 3.25 of the Patents
Regulations (Australia Statutory Rules 1991 No. 71), samples of materials
deposited
in accordance with the Budapest Treaty in relation to this Patent Request are
only to
be provided before: the patent is granted on the application; or the
application has
lapsed or been withdrawn or refused; to a person who is: a skilled addressee
without
an interest in the invention; and nominated by a person who makes a request
for the
furnishing of those samples.
WO 96/09381 PCT/FI95/00520
22
SEQUENCE LISTING
(1) GENERAL INFORMATION:
(i) APPLICANT: Helsinki University Licensing Ltd Oy
(ii) TITLE OF INVENTION: Promoter for the Receptor Tyrosine Kinase, TIE
(iii) NUMBER OF SEQUENCES: 9
(iv) CORRESPONDENCE ADDRESS:
(A) ADDRESSEE: Oy Jalo Ant-Wuorinen Ab
(B) STREET: Iso Roobertinkatu 4-6 A
(C) CITY: Helsinki
(E) COUNTRY: Finland
(F) ZIP: 00120
(v) COMPUTER READABLE FORM:
(A) MEDIUM TYPE: Floppy disk
(B) COMPUTER: IBM PC compatible
(C) OPERATING SYSTEM: PC-DOS/MS-DOS
(D) SOFTWARE: PatentIn Release #1.0, Version #1.25
(viii) ATTORNEY/AGENT INFORMATION:
(A) NAME: Karvinen, Leena
(C) REFERENCE/DOCKET NUMBER: 28203
(ix) TELECOMMUNICATION INFORMATION:
(A) TELEPHONE: +358 0 648606
(B) TELEFAX: +358 0 640575
(C) TELEX: 123505 jalo sf
(2) INFORMATION FOR SEQ ID NO:1:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 882 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: DNA (genomic)
(xi) SEQUENCE DESCRIPTION: SEQ ID NO:1:
TCTTAAGACATGCAACTCGTCTACGGCTATACCACTCTGAACGCGCCCGATCTCGGAAGA 60
CATGCAACTCAAATGTAAATACAGTAGAATATTACTTAGGTAGAAACTCCTGGTGATTTT 120
AAAAGATTGGAAAAGAATATGAGGAAGAGTTGAATAATGCAAATTCTAGTGTGTGTGCTA 180
CCGAAGTGAACACTTAATGCACAGTCTACAGACTAGGACATTTTATCGTGTGTTGTAAAA 240
TTGGGTAGAAACTTGTGTTTGTGAAAACTGAGCATTAAAACCTTACAGAGACCGTTTCTT 300
GTTTACTTTTG1~~~P.AAAAAP.AGAGTCACGTGAGCCTCATTTTGTATTTGTGTGTGTGTGT 360
GTGTGTGTGTCTCCCCTCCTCCCAGCGTGTGTGTGCTGGGAGGAGGGGAGACCCCAGAAC 420
AATGTCCTGCCTCCAAACCTTCTCAATAGGCGGAACGACTGGCTTCTCCCTTTCCTGTCT 480
SUBSTITUTE SHEET (RULE 26j
WO 96/09381 PCT/FI95/00520
23
CCCGTGCTCCAGCAATGCAGATGGAAGGGACCGAAGGGATGGGAGAGAGAGCCCAACCAT 540
CCCCAGATCTGTCCTTGTCACAACCTGCCTCCCACCTCTAATGCCCCCCCTTCCAGAGAC 600
TTCCAGGCCACACCCATCCCGGGCTTGTGGGGGCTGGACACGGGAGGACTACAGGCGACA 660
ACTCTTCCCACCCTCTCTCCCTGCCACCCCTCCTACCCTAACCATCATTTCCTCTTCCTC 720
CCCAGCACCGAGGTGCACTGAGCTGGACAGGCTGAACACTCAGACCCACAGCAACTGACC 780
CCGGGCCCAGCTGGCCTTGGCTGGCCCAGGGCAGCTTCCAGAGTATGGTCTGGTGGGGAT 840
CCTCTTTGCTGCTCCCCACTCTTTTCTTGGCCTCTCATGTTG 882
(2) INFORMATION FOR SEQ ID N0:2:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 935 base
pairs
(B) TYPE: nucleic
acid
(C) STRANDEDNESS:
single
(D) TOPOLOGY:
linear
(ii) MOLECULE
TYPE: DNA (genomic)
(xi) SEQUENCE
DESCRIPTION:
SEQ ID N0:2:
CGGCAAAATG AATGACACCTGGCAGACAATAAGCTGAAGCTTTCATTAGC AGCTTAAGCT60
GAGGACTATC TATGCAACCGATACTCCCTGTGTGCTCCCCGGGATGGTTA ATGTGAGGCC120
TTGTGGAGCG ATTGGCACCAAGGAAAGGAAGGACTAAGTCAGAAGTTCAA GTCCCAGCCT180
TGCCACAGCC TCAGGGTGCCCTCGAGCACAGCAAGCCTCAGTTTTCCCAT CTGTACAATG240
AGAGAGGTAC ACAAGGTAGACTCGAAGGCTCTTTGTTGCCAGGGCCCTGT GTTCCTTTGA300
GTGTATGTGC TTCTCAGGCCCACAGAGGTCCTTTGTGTTTCGTATGTGAA CTGCTCTCTA360
GGAAACCCAT GTAACTGTCTGTGTCCTGGGGCACATACATGAGGACTCAT GTGGGCCGTA420
TTGTGTGTTT GTGCCGGGGGGAGGGGAGACCCCAGAACAATGTCCCCCAC CCCACCCCCC480
TCCTCAATAG GCGAAGCGCACTGGCTTCCTCCCTTTCCTGCCTCCTGCCT CCTTTGTGCC540
AGCAAGACTG AGTACTGGAGGGAGACAGGGGATGGGAAAAATCAGTCCAG CTGTCCCCAG600
GTCTGCCCTT ACCATAACCTTCCCCCCACCTCAAGTGACTCCTCCCAGGC CACACCCATC660
CCCAGCCTTG TGGGGGCCAGATTGGGGGGCCTAGAGGCTCAAAGGCAGAA TGAGTCCTCC720
CACCCCCTAC CCTGCCACCCCTCCCACCCAAGCCACCTCATTTCCTCTTC CTCCCCAGCA780
CCGACCCACA CTGACCAACACAGGCTGAGCAGTCAGGCCCACAGCATCTG ACCCCAGGCC840
CAGCTCGTCC TGGCTGGCCTGGGTCGGCCTCTGGAGTATGGTCTGGCGGG TGCCCCCTTT900
CTTGCTCCCC ATCCTCTTCTTGGCTTCTCATGTGG 935
(2) INFORMATION FOR SEQ ID N0:3:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 131 amino acids
(B) TYPE: amino acid
SUBSTITUTE S4iEET (RULE 2~
WO 96/09381 PGT/FI95/00520
24
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: protein
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:3:
Gly Cys Val Lys Asp Cys Pro Gly Cys Leu His Gly Gly Val Cys His
1 5 10 15
Asp His Asp Gly Cys Val Cys Pro Pro Gly Phe Thr Gly Thr Arg Cys
20 25 30
Glu Gln Ala Cys Arg Glu Gly Arg Phe Gly Gln Ser Cys Gln Glu Gln
35 40 45
Cys Pro Gly Thr Ala Gly Cys Arg Gly Leu Thr Phe Cys Leu Pro Asp
50 55 60
Pro Tyr Gly Cys Ser Cys Gly Ser Gly Trp Arg Gly Ser Gln Cys Gln
65 70 75 80
Glu Ala Cys Ala Pro Asp His Phe Gly Ala Asp Cys Arg Leu Gln Cys
85 90 95
Gln Cys Gln Asn Gly Gly Thr Cys Asp Arg Phe Ser Gly Cys Val Cys
100 105 110
Pro Ser Gly Trp His Gly Val His Cys Glu Lys Ser Asp Arg Ile Pro
115 120 125
Gln Ile Leu
130
(2) INFORMATION FOR SEQ ID N0:4:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 15 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:4:
CCCACATGAG AAGCC 15
(2) INFORMATION FOR SEQ ID N0:5:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 32 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
SUBSTITUTE SHEET (RULE 26)
WO 96109381 PCT/FI95/00520
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:5:
TGAGATCTTG GAGTATGGTC TGGCGGGTGC CC 32
(2) INFORMATION FOR SEQ ID N0:6:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 21 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:6:
CTATTGAGAA GGTTTGGAGG C 21
(2) INFORMATION FOR SEQ ID N0:7:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 19 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:7:
GCTCTAGAAC TAGTGGATC 19
(2) INFORMATION FOR SEQ ID N0:8:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 20 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:8:
GAGACAGGGG ATGGGAAAAA 20
(2) INFORMATION FOR SEQ ID N0:9:
(i) SEQUENCE CHARACTERISTICS:
(A) LENGTH: 20 base pairs
(B) TYPE: nucleic acid
(C) STRANDEDNESS: single
(D) TOPOLOGY: linear
(ii) MOLECULE TYPE: cDNA
(xi) SEQUENCE DESCRIPTION: SEQ ID N0:9:
GAAGATCGCA CTCCAGCCAG 20
SUBSTITUTE SHEET (RULE 26)