Note: Descriptions are shown in the official language in which they were submitted.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
DNA sequences for enzymatic synthesis of polyketide or
heteropolyketide compounds
The present invention relates to DNA sequences for enzy-
matic synthesis of polyketide or heteropolyketide compounds
produced by the bacterium Sorangium ce11u1osum.
Background and introduction
This patent application describes DNA sequences for the
enzymatic synthesis of polyketide and/or heteropolyketide
structures synthesized by the myxobacterium Sorangium cellulo-
sum. Several of these compounds have known cytotoxic, immuno-
suppressive, antibiotic and fungicidal biological activity,
with the epothilones having been most studied and character-
ized. The fermentation of large quantities of secondary me-
tabolites from microorganisms, especially from myxobacteria, is
a time consuming and difficult process that often involves com-
plications (e. g. contamination, low product yield, difficult
isolation and purification). Therefore it would be advanta-
geous to use a well-characterized organism for such fermenta-
tions. After cloning of the desired biosynthetic genes one
could create such an organism via genetic engineering and ma-
nipulate the biosynthesis of the compound. Identified sequences
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
2
can be cloned into optimized expression vectors and generate
recombinant cell lines that overproduce polyketide structures.
Polyketide synthases (PKS) and non-ribosomal peptide syn
thetases (NRPS) represent macromolecular and multifunctional
enzymes which are characterized by a modular architecture. PKS
condenses activated carbonic acids (usually acetate and propi-
onate) and reduce the resulting 2-keto acid intermediates step-
wise in a fatty acid biosynthesis-like fashion. Responsible for
each reaction step is a specific domain that recognizes, acti-
vates, condenses and reduces the carbonic acid. Depending on
the presence of these domains in the corresponding modules,
every reduction stage can occur in the final product (Rawlings,
Nat. Prod. Reports 14, 523-556 [1997]; for a review, see Chem.
Rev. ~7, 2463-2760 [1997]). A typical example for the biosyn-
thesis of a polyketide is the macrolide antibiotic erythromycin
(Staunton and Wilkinson, Chem. Rev. 97, 2611-2630 [1997]).
NRPSs are also modular enzymes and condense via peptide bonds
amino acids to low molecular weight bioactive substances like
bacitracin or tyrocidin. Typical domains of these systems acti-
vate the amino acid and condense it with the growing peptide
chain. Methylations, epimerisations and modifications via addi-
tional protein domains are possible (Stachelhaus and Marahiel,
FEMS Microbiol Lett. 125, 3-14 [1995]). Both types of enzymes
(NRPS and PKS) share the modular organization of the proteins
in which specific catalytic domains are responsible for recog-
nition, activation, condensation and modification of the single
elongation units . The growing chain of amino acids and/or car-
bonic acids is extended through the action of one module adding
one unit. The domains of each module carry the active centers
responsible for the enzymatic steps of the biosynthesis.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
3
Little is known about the biosynthesis of biologically ac-
tive polyketides and polypeptides from myxobacteria. Fragments
of the biosynthetic gene clusters of soraphen and saframycin
have been described from Sorangium cellulosum So ce26 and Myxo-
coccus xanthus, respectively (Schupp et al., J. Bacteriol. 177,
3673-3679 [1995] and Pospiech et al., Microbiology 141, 1793-
1803 [1995]). We have constructed genomic libraries of the
epothilone producer Sorangium cellulosum So ce90. Gene probes
based on PKS and PS genes were used to isolate recombinant cos-
mids, which were then sequenced and characterized. Several
unique pathways containing PKS, PS, or a combination of both
types of genes were identified, demonstrating that this organ-
ism is potentially a rich source of novel bioactive compounds.
A subject of the present invention is therefore to provide
DNA sequences according to claim 1 the expression products of
which perform or are involved in the enzymatic biosynthesis,
mutasynthesis or partial synthesis of polyketide or hetero
polyketide compounds. The DNA sequences may be inserted into
well known and optimized expression vectors by commmon tech
niques of molecular biology, thus allowing transformation, se-
lection and cloning of cells, which cells are then capable of
synthezising polyketide or heteropolyketide compounds by fer-
mentation. Using an overproducing clone allows the desired
polyketide or heteropolyketide compounds be easily produced and
recovered in high amounts. Further, knowledge of the localiza-
tion of regulatory DNA segments and individual structural genes
allows "site-directed mutagenesis" using common techniques for
genetic engineering, ar_d thus construction of optimized enzymes
("protein engineering"for fermentative synthesis of polyketi-
de or heteropolyketide compounds.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
4
The inventicr. thus further relates to a recombinant ex-
pression vector according to claim 16, cells transformed there-
with according tc claim 17 and to a process for enzymatic bio-
synthesis, mutasyr_thesis or partial synthesis of polyketide or
heteropolyketide compounds according to claim 23.
Preferred and/or advantageous embodiments of the present
invention are subject-matter of the subclaims.
In brief, the invention consists of (1) cloned Sorangium
cellulosum polyketide synthase (PKS) and/or peptide synthetase
(PS) biosynthetic cluster DNA and (2) the nucleotide sequence
and predicted protein coding sequences of the cloned DNA. The
invention can be used for, but not limited to, (a) increasing
yields of PKS product in Sorangium cellulosum (e. g., by ampli-
fication or genetic modification of the epothilone gene cluster
or its component parts), (b) increasing yields of polyketide
and/or peptide synthetase product in a heterologous system by
transfer of the corresponding gene cluster or its component
parts, which may be followed by amplification or genetic modi-
fication of the PKS and/or PS gene cluster or its component
parts, (c) modification of the polyketide and/or peptide syn-
thetase product chemical structure in either Sorangium ce11u1o-
sum or a heterologous host (e.g., by genetic modification of
the corresponding gene cluster or its component parts) and (d)
for the detection of genes and gene products involved in making
polyketides or related molecules in other organisms (e.g., by
hybridization or complementation assays). DNA sequence and
analysis is presented for the following cosmids and plasmids:
- A2 cosmid as defined in claim 6
- the pEPOcos6 region (overlapping of pEPOcos6 and pEPOcos7)
as defined ir_ claim 7
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
- pEPOcosB cosmid as defined in claim 10
- A5 cosmid as defined in claim 12
- Sau4 (10 kb plasmid) as defined in claim 14
5 The invention is now described in more detail by examples and
for illustration only. The examples are not to be construed as
any limitation of the scope.
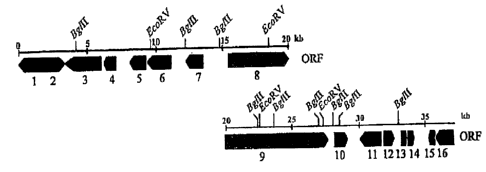
Figure 1 is a restriction map of one of the DNA sequences of
the present invention (cosmid A2 insert) indicating also the
localization of regulatory DNA segments and the individual
structural genes ("open reading frames" or ORFs) 1 to 16.
Figure 2 shows the open reading frames found on pEPOcos6 region
DNA sequence data from A2 cosmid are as defined in claim 6.
Table 1 correlates ORFs 1 to 16 found on A2 cosmid with the re-
spective biological function (Regulators, Enzymes).
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
Table 1
gene/function position
ORF 1 regulatory element 1666 - 1
ORF 2 regulatory element 1605 - 3338
ORF 3 acyl-t-RNA synthetase 6100 - 3398
ORF 4 monooxygenase 7110 - 6374
ORF 5 amino transferase 9590 - 8433
ORF 6 L-dopa decarboxylase 11393 9855
-
ORF 7 oxidoreductase 13656 12712
-
ORF 8 polyketide synthase 15374 18984
-
ORF 9 polypeptide synthetase 20003 27889
-
ORF 10 peptidase 28251 29402
-
ORF 11 regulatory element 31720 30401
-
ORF 12 sigma factor 31982 32932
-
ORF 13 regulatory element 33128 33613
-
ORF 14 regulatory element 33661 34007
-
ORF 15 transcr=ption regulator 35611 35255
-
ORF 16 signal ~ransduction 37856 35730
-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
7
Working Examples
A. Construction of a Sorangium cellulosum cosmid library
1. Isolation of genomic DNA from S. ce11u1osum So ce90
a. Sorangium cellulosum So ce90 was spread onto solid CA-2
agar and incubated at 3G°C for 5-7 days. CA-2 agar is prepared
by autoclaving 18 g Bacto-agar (Difco Laboratories, Detroit,
MI) in 800 ml dHzO for 20 min at 121°C and cooling to 50-
55°C in
a water bath. The following filter-sterilized solutions are
added to the agar: 20% (w/v) glucose, 50 ml; Solution A (7.5%
[w/v] KN03,7.5% K2HP04), 10 ml; Solution B (1.5% [w/v]
MgSG'7H20) , 10 ml; Solution C (0.2% [w/v] CaCl2'2H20, 0.15% [w/v]
FeCl3), 10 ml; 1 M HC1, 1 ml; autoclaved 4-day old Sorangium
cellulosum broth, 100 ml. A sample of cells was removed from
the plates with a sterile loop and inoculated into 50 ml of
G5lt medium in a 250 ml Erlenmeyer flask. G5lt consists of 0.5%
starch (Cerestar), 0.2% tryptone, 0.1% yeast extract, 0.05%
CaC-.:~, 0.05% MgSO; 7H20, 1.2% 4- (2-hydroxyethyl) -1-piperazine
etranesulfonic acid (HEPES), 0.2% glucose, pH 7.6. The flasks
were shaken at 30°C, 160 rpm until a dense orange bacterial
growth was obtained (cG. 5-7 d.). The cells were pelleted by
centrifugation at 6,000 x g and used immediately or stored fro
zen at -20°C.
The protocol used nor isolating chromosomal DNA from bac-
ter-a using hexadecyltrimethylammmonium bromide (CTAB) has been
described previously (A~.subel et al., Current Protocols in Mo-
lec~;:lar Biology, John Y1;_iey and Sons, New York, 1990). The pre-
cio-sated DNA was recovered with a bent Pasteur pipette, washed
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
8
with 70% and 95% ethanol, air-dried, and resuspended in 0.5 ml
TE buffer (0.01 M Tris-HC1, 0.001 M ethylenediaminotetraacetic
acid [EDTA] , pH 8 . 0 ) .
b. Alternatively, genomic DNA was isolated from S. cellulosum
cells cultured as described in section A.1 using the Midi
Qiagen Blood & Cell Culture DNA purification Kit (Qiagen,
Hilden, Germany) following the Qiagen Genomic DNA Handbook pro-
tocol for bacterial DNA isolation (1997, Qiagen, Hilden, Ger-
many, p. 29 ff.). In order to obtain high molecular weight
chromosomal DNA the precipitated DNA was recovered with a bent
pasteur pipette as described in section A.1.
2. Isolation of plasmid DNA
a. pFD666: pFD666 is a bifunctional E. coli-Streptomyces cosmid
cloning vector (see Denis and Brzezinski, Gene 111, 115-118
[1992]). To maintain stability of large inserts, it is present
in low-medium copy number when replicated in E. coli. For this
reason, isolation of sufficient pure DNA to carry out cloning
experiments was difficult using commercial kits with standard
protocols. A modified procedure was therefore used to obtain
pFD666 DNA. A 10 ml culture of DH10B(pFD666) was grown for 16-
20 hr at 37°C in LB (1% tryptone, 0.5% yeast extract, 0.5%
NaC~, pH 7.0) medium containing 50 ug/ml kanamycin sulfate.
Fif~y ml of LB + kanamycin was inoculated to a starting OD6oo of
ca. 0.25 and shaken at 300 rpm, 37°C, until the OD6oo reached
ca. 0.6. Five hundred ml of LB + kanamycin medium in a 2 1
flask was inoculated with 25 ml of this culture and incubated
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
9
under the same conditions for 2.5 hr. Chloramphenicol ( 2.5 ml
of a 34 mg/ml solution in 100% EtOH) was added and the incuba-
tion continued for an additional 16-20 hr. (The previous steps
were performed according to Maniatis et a:l. Molecular Cloning:
A Laboratory Manual, Cold Spring Harbor Laboratory, Cold Spring
Harbor, NY, 1989.) Cells were pelleted for 10 min, 16,000 x g .
They were resuspended in 9 ml of 50 mM glucose/25 mM Tris-HCl
(pH 8.0)/10 mM EDTA and transferred to a 50 ml disposable cen-
trifuge tube. One ml of a freshly-prepared 10 mg/ml lysozyme
solution in 10 mM Tris-HC1, pH 8.0 was added and the cell sus-
pension incubated in a 37°C water bath for 10 min. Twenty ml of
a freshly-prepared 0.2 NaOH/1% sodium dodecyl sulfate (SDS) so-
lution was added and the tube inverted gently 5-7 times to mix
the contents. After 5 min at room temperature, 15 ml of 5 M po-
tassium actate (pH 4.8) was added and the tube inverted sharply
3-4 times. The tube was centrifuged at 6,000 x g for 10 min at
4°C and the supernatan;_. poured though 2 layers of sterile
cheese cloth into a fresh 50 ml disposable tube. Isopropanol to
a final concentration of 0.6% was added and the contents of the
tube mixed several times. The precipitated nucleic acid was
centrifuged at 6,000 x g for 10 min at 4°C. The pellet was
washed with 70% EtOH and any excess EtOH was aspirated from the
pellet, which was allowed to air dry for 5 min. It was resus-
pended in 5 ml of 50 mM 3-(N-Morpholino)propanesulfonic acid
(MOPS)/75o mM NaCl, pH 7.0 and added to an equilibrated to
QIAfilter Midi column (Qiagen, Chatsworth, CA). The manufac-
turer's protocol for washing and eluting the plasmid DNA was
followed.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
b. SuperCos: SuperCos -lasmid DNA was purchased from Strata-
gene (La Jolla, CA) .
3. Preparation of ca. 38-47 kb Sau3Al fragments of S. ce11u1o-
5 sum chromosomal DNA
a. S. cellulosum chromosomal DNA prepared as described in sec-
tion A.l.a was partially- cleaved with restriction endonuclease
Sau3A1 in a 1000 ~l reac~ion volume consisting of 50 ~,g chromo-
10 somal DNA, 5 units enzv~r:e (Promega, Madison , WI), 0.006 M
Tris-HCl, 0.006 M MgCl~, 0.10 M NaCl, and 0.001 M dithiothrei-
tol (pH 7.5) for 5 min at 37°C. The reaction mixture was ex-
tracted once with an eaual volume of 1:1 phenol: chloroform.
After centrifugation, tre upper aqueous phase was saved, to
which 0.1 vol. of 3 M sodium acetate and 0.6 vol. isopropanol
was added. DNA was pe_leted by centrifugation for 5 min at
16,000 x g in a microfua~ and washed once with 0.5 ml 70% EtOH.
After drying in a SpeeaVac (Savant Instruments, Farmingdale,
NY) for 5 min, the pell~_ was resuspended in 0.1 ml TE buffer.
The DNA was layered oncco of a 12 mi 10-40% sucrose gradient
prepared in TE buffer aceritrifuged at 113,600 x g for 16 hr,
10°C using a Beckman SW40Ti rotor (Beckman Instruments, Palo
Alto, CA). Five hundrea ~1 aliquots of the gradient were re-
moved using a pipetor beginning at the top of the tube. Samples
(5 ~1) of the fracti:Ts were analyzed by electrophoresis
through a 0.5% agarose ~~l in TAE buffer (0.04 M Trizma base,
0.02 M acetic acid, any: 0.001 M EDTA, pH 8.3) containing 0.5
~.g/ml ethidium bromide -cr 6 hr at 100 V. Fractions containing
DNA fragments of ca. 4G-5 kb were identified by comparison to
a high molecular wei_=~ DNA standard (Life Technologies,
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
11
Gaithersburg, MD). Sucrose was diluted from the corresponding
0.5 ml fraction by addition of 0.5 vol. TE. Subsequently, DNA
was precipitated by addition of 0.1 vol. 3 M sodium acetate and
0.6 vol. isopropanol. DNA was pelleted by centrifugation at
16, 000 x g for 10 min in a microfuge. DNA was washed with 0.5
ml 70% EtOH and dried in a SpeedVac with moderate heat for 10
min. Finally, the DNA was resuspended in distilled H20 at a
concentration of 0.5 mg/ml.
b. Alternatively, 10 ~.g of S. cellulosum chromosomal DNA pre-
pared as described in A.l.b was treated with 0.3 U Sau3A1 (New
England Biolabs, Beverly, MA) for 1 h at 37°C in 400 ~,l of the
supplier's recommended reaction buffer. Formation of DNA frag-
ments of about 40 kb in size was checked by comparison of the
motility behavior with high molecular weight DNA standards af-
ter a 0.3% agarose gel electrophoresis. An equal volume of phe-
nol:chloroform (1:1) was added, mixed and centrifuged. The up-
per aqueous phase was recovered and 0.1 vol. of 3 M sodium ace-
tate and 0.6 vol. of isopropanol were added. After centrifuga-
tion, the precipitated DNA was washed twice with 0.5 ml 70% ice
cola ethanol and finally air-dried. The DNA fragments were re-
suspended in 100 ~1 shrimp alkaline phosphatase reaction buffer
and dephosphorylated for 150 min. at 37°C using 2 U shrimp al-
kaline phosphatase (Amersham Life Science, Cleveland, OH). A
phenol:chloroform extraction followed as described above. Fi-
nally, the DNA was precipitated by addition of 0.1 vol. 3 M so-
diu~. acetate and 0.6 vol. isopropanol, dried, and dissolved in
TE ::,uf fer .
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
12
4. Preparation of cosmid libraries
a. Using pFD666: Vector pFD666 was cleaved with restriction
endonuclease BamHI in a 0.02 ml reaction volume consisting of 2
~.g plasmid DNA, 10 units of BamHI (Promega) , 0.006 M Tris-HC1,
0.006 M MgCl2, 0.05 M NaCl, and 0.001 M dithiothreitol (pH 7.5)
for 90 min at 37°C. Five ~.l of lOx alkaline phosphatase buffer
(0.5 M Tris-HCl [pH 9.3], 0.01 M MgClz, 0.001 M ZnCl2, 0.01 M
spermidine) was added to the reaction followed by alkaline
phosphatase (0.01 units/pmol ends; Promega) and distilled H20
to a final volume of 0.05 ml. The sample was incubated for 30
min at 37°C and a second aliquot of phosphatase was added. Af-
ter a further 30 min at 37°C, 0.3 ml of stop buffer (0.01 M
Tris-HCl [pH 7.5], 0.001 M EDTA, 0.2 M NaCl, 0.5% SDS) and 0.35
ml of 1:1 phenol; CHC13 was added to the reaction. The sample
was mixed gently several times by inversion and centrifuged at
16,000 x g for 3 min to separate the phases. The aqueous layer
was removed to a new microfuge tube . 0 . 1 vol . 3 M sodium ace-
tate and 2 vol. 100% EtOH were added and the precipitated DNA
pelleted by centrifugation at 16,000 x g for 10 min. Liquid was
removed by aspiration and the pellet washed once with 0.5 ml
70% EtOH. The DNA was dried in a SpeedVac and resuspended in TE
buffer to 0.5 mg/ml.
Digested, phosphatase-treated pFD666 was ligated to the
partially-cleaved chromosomal DNA (see sections A.3.a and
B.l.a) in a 0.005 ml reaction consisting of 1 ~.g pFD666, 1 ~,g
S. cellulosum DNA, 0.03 M Tris-HCl (pH 7.8}, 0.01 M MgCl2, 0.01
M dithiothreitol, and 0.0005 M adenosine-5'-triphosphate and
1.5 Vdeiss units of T4 DNA ligase (Promega}. The reaction was
carried out at room temperature for 2 hr. The entire reaction
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
13
mix was packaged into bacteriophage 7~ in vitro using Packagene
extracts (Promega) according to the manufacturer's directions.
The entire packaging reaction (0.5 ml) was diluted with 4.5 ml
SM buffer (per liter: 5.8 g NaCl, 2 g MgS04.7Hz0, 1 M Tris-
HCl [pH 7.5] , 5 ml 2% ge~.atin solution) . Transfection was per-
formed by adding 10 ml c. an overnight culture of E. coli DHSa
that had been grown in LB medium with 0.01 M MgS04 and 0.2%
maltose to the diluted p_-gage and incubating at 37°C for 20 min.
0.8 ml of LB was added a::d the cells shaken at 225 rpm for 1 hr
at 37°C. Cells were pel_eted, resuspended in LB, and spread
onto a 150 mm LB + kana:«ycin agar plate. After 3 d. at 30°C,
the colonies were harvested by picking ca. 800 colonies into
2.0 ml LB + kanamycin medium containing 20% glycerol, freezing
on dry ice, and storing at -70°C. In addition, six kanamycin-
resistant colonies were inoculated into 2 ml LB + kanamycin
liquid medium and incubated at 37°C, 250 rpm, for 18-24 hr.
Cosmid DNA was prepared casing a standard alkaline lysis proce-
dure starting with 1.5 rr.-~ of the culture. DNA was digested with
restriction endonuclease PstI and samples electrophoresed on a
0.8% TAE agarose gel for 1.5 hr at 100 V. A unique restriction
pattern was noted in eac_~ sample and the total size of the in-
sert was calculated to be between 40 and 45 kilobases.
b. Using SuperCos: 30 ~g of vector SuperCos was digested with
XbaI (New England Biolabs, Beverly, MA) for 210 min at 37°C in
100 ~1 of the recommended reaction buffer. Ten ~,l sodium ace-
tate and 60 ~1 isopropa_:ol was added before the solution was
centrifuged for 30 min w_ 16,000 x g. The precipitated DNA was
washed twice with 500 ~:= ice cold 70 % ethanol . The vector DNA
was precipitated and ai=-dried, dissolved in 135 ~l shrimp al-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
14
kaline phosphatase reaction buffer and treated with 2.5 U
shrimp alkaline phosphatase for 150 min. After heat inactiva-
tion of the enzyme at 75°C for 20 min, a phenol:chloroform ex-
traction was performed as described in section 1. c. The DNA,
resuspended in 100 ~.l BamHI restriction buffer was hydrolyzed
with 15 U BamHI (New England Biolabs, Beverly, MA) for 180 min.
A phenol:chloroform extraction followed (see section A.3). The
SuperCos DNA was precipitated by additon of 0.1 vol 3 M sodium
acetate and 0.6 vol isopropanol, centrifuged at 16,000 x g, and
resuspended in 50 ~.1 TE buffer.
Four ~.g of digested vector DNA was ligated with 10 ~.g par-
tially hydrolyzed genomic DNA from S. cellulosum (as described
in section A.3.b) in a final volume of 20 ~,1 using 2 U T4 DNA
ligase and the appropriate reaction buffer (Gibco BRL, Eggen-
stein, Germany). The reaction was carried out at 16°C over-
night. The reaction mixture was packaged into phage particles
using the Gigapack III XL packaging extract kit (Stratagene)
according to the manufacture's protocol. Treatment of packaging
reaction mixture and transfection of E, coli SURE (Stratagene)
was performed as described in 4.a. Transfected cells were con-
centrated by centrifugation, resuspended in fresh LB medium and
distributed on LB agar plates containing 50 ~,g/ml-lkanamycin.
The plates were incubated overnight at 30°C. 1600 recombinant
clones were transferred into 96 well microtiter plates filled
with 80 ~.l LB medium containing 50 ~,g/ml~kanamycin per well and
propagated overnight at 30°C. The following day the microtiter
plates were used to inoc~.~late a second set of microtiter plates
in order to obtain a duplicate of the recombinant clones. Each
well of the original set of microtiter plates was supplemented
with 80 ~l 50 o glycero~. and the entire plate stored at -70°C.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
20 randomly chosen transformants were inoculated into 3 ml
LB medium with 50 ug/ml-1 kanamycin and incubated over night at
37°C in order to isolate plasmid DNA using the Qiagen plasmid
extraction kit (Qiagen, Hilden, Germany). Restriction fragment
5 analysis of the recombinant cosmids using the restriction endo-
nucleases PstI and BglII indicated that the cosmids contained
inserts of approximately 35 to 42 kb in size.
B. Construction of a S. cellulosum plasmid library
1. Preparation of 8-12 kb fragments of S. ce11u1osum chromoso-
mal DNA.
S. cellulosum chromosomal DNA prepared as described in sec-
tion A.l.a was partially cleaved with restriction endonuclease
Sau3Al in a 100 ~tL reaction volume consisting of 5 ~g chromoso-
mal DNA, 5 units enzyme {Promega, Madison , WI), 0.006 M Tris-
HC1, 0.006 M MgCl2, 0.10 M NaCl, and 0.001 M dithiothreitol (pH
7.5) for 4 min at 37°C. The digested DNA was electrophoresed
through a 11 x 14 cm 0.8% TAE-agarose gel for 18 hr at 17 V.
Fragments of 8 -12 kb were cut f rom the gel and purl f ied using
the QIAquick Gel Extraction Kit using the manufacturer's proto-
col (Qiagen).
2. Preparation of the plasmid library
?lasmid pZero2.1 (Invitrogen, Carlsbad, CA) was cleaved with
res~riction endonuclease BamHI in a 0.02 ml reaction volume
consisting of 1 ~g plasmid DNA, 10 units of BamHI (Promega),
O.G06 M Tris-HC1, 0.006 M MgClz, 0.05 M NaCl, and 0.001 M di-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
16
thiothreitol ( pH 7.5) for 20 min at 37°C. 0.08 ml of dH20 and
0.1 ml of 1:1 phenol:CHCl3 was added. The sample was briefly
vortexed and centrifuged at 16,000 x g for 2 min. The aqueous
layer was removed to a new microfuge tube. 0.1 vol. 3 M sodium
acetate and 2 vol. 100% EtOH were added and the precipitated
DNA pelleted by centrifugation at 16,000 x g for 10 min. Liquid
was removed by aspiration and the pellet washed once with 0.5
ml 70% EtOH. The DNA was dried in a SpeedVac and resuspended in
TE buffer to 0.004 ~g/ml. Digested pZero2.1 was ligated to the
partially-cleaved chromosomal DNA in a 0.01 ml reaction con-
sisting of 0.004 ~.g pZero2.l, 0.05 ~g S. cellulosum DNA, 0.03 M
Tris-HCl (pH 7.8), 0.01 M MgCl2, 0.01 M dithiothreitol, and
0.0005 M adenosine-5'-triphosphate and 1.5 Weiss units of T4
DNA ligase (Promega). The reaction was carried out at room tem-
perature for 2 hr. 0.015 ml dH20 and 0.25 ml of 1-butanol were
added, the sample vortexed briefly, and centrifuged at 16,000 x
g for 10 min. Liquid was aspirated away from the pellet and the
sample dried in a Speedvac for 5 min. The ligated DNA was re-
suspended in 0.005 ml dHzO and mixed with 0.04 ml of electro-
competent Escherichia coli DH10B cells (GIBCO/BRL, Gaithers-
burg, MD). The sample was placed into a pre-chilled 0.2 mm-gap
electroporation cuvette and transformed into the bacteria by
electroporation using a BioRad Gene Pulser II unit (BioRad,
Hercules, CA) at 25 ~F and 200 S2. 0.96 ml SOC medium (0.5%
yeast extract, 2% tryptone, 10 mM NaCl, 2.5 mM KC1, 10 mM
MgCl2, 20 mM MgS04, 20 mM glucose) was mixed with the cells and
transferred to a 1.5 ml microfuge tube. The sample was incu-
bated at 37°C, 225 rpm, for 1 hr. Aliquots of the cells were
spread onto an LB agar + kanamycin and incubated at 37°C for 20
hr to estimate the number of transformants obtained. Six kana-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
17
mycin resistant colonies ~~:ere confirmed to contain an insert of
the expected size as described in section A.4.a.
C. Identification of cosmids possessing polyketide synthase
genes
1. Colony blot hybridizations using cosmid library in pFD565:
A 20 x 20 cm sheep of Duralon UV membrane (Stratagene)
was placed on top of a 2..5 x 24.5 cm square bioassay dish con
taining 250 ml LB agar kanamycin. An aliquot of the frozen
cosmid library in 1 ml L? medium was spread on the filter. The
plate was incubated at 37VC for 24 hr. Colonies were replicated
onto two fresh filters which were placed onto LB + kanamycin
agar medium and incubates at 28°C for 18 hr. Lysis of cells and
neutralization of released DNA was performed according to di-
rections that were pro-fided with the filters. The DNA was
crosslinked to the filters using a W Stratalinker 2400 unit
(Stratagene) in the auto crosslink mode. Cell debris was re-
moved by placing the filers in a container with a solution of
3 X SSC (20 X SSC contai_~s, per liter, 173.5 g NaCl, 88.2 g so-
dium citrate, pH adjuste~ to 7.0 with 10 N NaOH), O.lo SDS and
rubbing the lysed colonies with a Kimwipe. The filters were
then incubated at least 3 hr with the same wash solution for at
least 3 hr at 65°C. The plasmid library was treated similarly
except cells were spread onto a 137 mm circular Duralon UV mem-
brane placed on top of .~ 150 mm petri dish containing 80 ml LB
agar + kanamycin.
For hybridizations, G probe consisting of a 650-base pair
(bp) polymerase chain (=C~) fragment representing a portion of
a S. cellulosum polyketie synthase gene was used. The fragment
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
18
was amplified using primers to consensus regions of Type I
(macrolide) polyketide synthase (PKS) genes (Swan et al., Mol.
Gen. Genetics 242, 358-362. [1994]). A series of sense and anti
sense oligonucleotides were prepared for PCR studies as indi
Gated in the following table 2:
Table 2
Oligo- I. DNA sequer_cP (5'-~ 3'~ Corresponding
nucleotide amino acid
(sequence
120 CGGT(C/G)AAGTC(C/G)AACATCGG KSNIGHT
(sense)
121 (anti- GC (A/G) ATCTC (A/G) CCCTGCGA (A/G) HSQGEIA
TG
sense)
122 I GT (C/G) GACAC (!./G) GC (C/G) TGCTCVDTACSS
i (C/G)
(sense)
123 GG (C/G) AC (C/G) AACGC (C/G) CACGT GTNAHVI
(C/G) A
(sense) T
12~. (anti-CCCTG(C/G)CC;C/G)GGGAA(C/G)ACGAA FVFPGQG
~
sense)
The selection of C or G where necessary in t:he third position
of a codon reflects the very high overall G + C content of S.
cellulosum (ca. 700). Conditions for PCR were as follows: 0.01
M T=is-HCl (pH 9.0) , 0.;~5 M KCl, 0.003 M MgC:lz, 0.1% Triton X-
10:,.200 ~M of each primer, 2.5 U Taq DNA polymerase (Promega),
5.0% dimethyl sulfoxide ;Sigma), and 0.01 ~g of S. cellulosum
chromosomal DNA in a 0.;5 ml reaction volume. Reactions were
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
19
carried out in a Perkin-Elmer Model 480 Thermocycler (Perkin-
Elmer Corporation, Foster City, CA) under the following condi-
tions : 94°C, 1 min; 50°C, 1 min, 72°C, 1 . 5 min for a
total of 30
cycles. Each possible combination of sense and anti-sense prim-
ers were tried. A 650-by and 350-by fragment was amplified us-
ing oligos 120 + 124 and 123 +124, respectively. The sequence
of the fragments were determined using the ALFexpress AutoRead
kit to fluorescently label the DNA, which was analyzed on an
ALFexpress sequencing apparatus (Pharmacia). The data indicated
both PCR fragments possessed significant homology to polyketide
synthase genes of Type I antibiotics. The 650-by fragment was
chosen for hybridization experiments.
The fragment was labeled with 3zP-dCTP using the NEBlot kit
(New England Biolabs, Beverly, MA) and purified on a Bio-Spin 6
column (BioRad, Hercules, CA.). Duplicate blots were pre
hybridized in 3 X SSC (1 X SSC contains 0.15 M sodium chloride
and 0.015 M sodium citrate, pH 7.0), 4 X Denhardt's solution
(100 X is 2% Ficoll [Type 400], 2% polyvinylpyrrolidone, and 2%
bovine serum albumin [Fraction V]), and 100 ~eg/ml sheared, de-
natured salmon sperm DNA; all reagents purchased from Sigma
Chemicals, St. Louis. The labeled DNA was heated in a boiling
water bath for 5 min to denature the strands, cooled on ice,
and added to the pre-hybridization solution. The filters were
incubated for at least 18 hr in a roller bottle hybridization
oven. They were transferred to new bottle, then washed two
times in 2 X SSC, 0.1% SDS at 70°C for 30 min (moderate strin-
gency) . The membranes were placed on Whatman 3MM paper to re-
mov~ excess liquid, covered with Saran Wrap, and exposed to
au~~radiography film (Kodak X-GMAT LS) with two intensifying
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
screens . The cassette was placed at -70°C and developed at ap-
propriate intervals.
Approximately 100 colonies were seen to have hybridized on
the duplicate filters. Fourteen of these were isolated from the
5 master plate and grown in 4 ml LB + kanamycin medium for 20-24
hr, 37°C, 250 rpm. Plasmid DNA was prepared using the standard
alkaline lysis method ar_d digested with restriction endonucle-
ase PstI. The digested DNA was electrophoresed on a 0.8% aga-
rose gel in TAE for 3 hr at 100 V. Fragments were transferred
10 to Duralon UV using the VacuGene XL vacuum blotting unit (Phar-
macia) and the recommended alkaline denaturation protocol. Hy-
bridization with radioac~ively-labeled PCR fragment and washing
were carried out as described above. Two prominent types of
cosmids were observed; one contained PstI fragments of ca. 7.0,
15 5.0, and 1.1 kb (pEPOcosS and pEPOcos7) that hybridized to the
probe; the other type had fragments of ca. 6.0 and 3.6 kb
(pEPOcos8 and pEPOcosl3i which were homologous to the probe.
Restriction analysis cor.~irmed that cosmids showing identical
hybridization patterns had identical or overlapping inserts.
20 PCR reactions using primers representing consensus sequences of
Type I PKS genes were performed using the isolated cosmid DNA
as template under condi~ions described above, except ca. 0.01
g of cosmid DNA was included as template. Cosmids pEPOcos6 and
pEPOcos8 amplified the 550-by fragment seen when oligonucleo
tides 120 + 124 were used, while pEPOcos8 and pEPOcosl3 sup
ported amplification of an 1100-by PCR fragment with oligos 122
and 124. The latter fragment was sequenced and confirmed to
possess strong similarity to Type I PKS genes. These data con-
firm that the recombinar_~ cosmids are related to each other and
that all contain PKS-like genes.
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
21
2. Colony blot hybridizations of plasmid library in pZero2.l:
A 137-mm circle o. Duralon UV membrane was placed on top
of a 150-mm containing 7J ml LB agar + kanamycin. An aliquot of
the plasmid library (representing ca. 2,000 recombinant colo
nies) in 0.5 ml LB medi~:~~ was spread on the filter. The plate
was incubated at 37°C fc= 20 hr. Colonies were replicated onto
two fresh filters whicr. were placed onto LB + kanamycin agar
medium and incubated at 37°C for 6 hr. The filters were proc-
essed for hybridization as described in Section C.1. Out of 8
positive colonies detected, one contained a plasmid with a DNA
region not encoded by e~=her pEPOcos6 or pEPOcos8. This plas-
mid, called Sau4, was characterized in more detail.
3. Colony blot hybridizations of cosmid library in SuperCos:
The recombinant E. coli clones from the microtiter plates
(see section 4. b) were used to produce two identical sets of
hybridization filters ir_ order to identify cosmids carrying PKS
and PS genes. The recorr~.~inant clones were spotted onto 2 sets
of 22 x 22 cm LB agar plates containing 50 ~.g/ml kanamycin.
Each plate contained 38~ T ones therefore representing 4 micro-
titer plates . The clones were incubated at 30°C overnight . Af -
ter pre-cooling for appr:~ximately 3 h at 4°C, 20 x 20 cm Hybond
N+ Nylon membranes (A-~:ersham, Braunschweig, Germany) were
placed onto the agar sur=aces. After 2 min. the membranes were
removed and placed for -~ min. on Whatman 3 MM paper (Whatman
paper Ltd., Maidstone, =.:gland) soaked with denaturation solu-
tion ( 0 . 5 N NaOH, 1 , 5 M waCl ) before they were transfered onto
Wha~man 3 MM paper sat~:=~ated with neutralization solution (1 M
Tris-HCI, pH 7.5, 1.5 I~ \TaCl) . Subsequently the membranes were
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
22
placed onto Whatman 3 MM paper soaked with 2 X SSC (0.3 M NaCl,
0.03 M sodium citrate, pH 7.2) for 10 min. The membranes were
baked for 40 min at 85°C. Then, each membrane was overlayed
with 5 ml Proteinase K solution (2 mg/ml Proteinase K in 2 x
SSC) and incubated at 37 °C for 90 min. Finally, cell debris
was removed by wiping the membranes with a Kimwipe pre-wetted
with 2 X SSC.
As we were seeking in particular to identify biosynthetic
pathways containing botr PKS and PS genes, the following hy-
bridization strategy was taken: The screening was initially fo-
cused on ketosynthase domains from type I PKSs and on the ade-
nylation domain from PSs . Target--specific primers were used to
amplify DNA fragments of the corresponding genes from chromoso-
mal DNA of S, cellulosum by PCR. The fragments obtained were
then cloned, sequenced and the deduced amino acid sequence com-
pared to known ketosynthase and adenylation domains of PKS and
PS, respectively. In a second step these PCR fragments were
used as gene probes to detect recombinant cosmids of the S.
cellulosum cosmid library.
Oligonucleotides based on conserved amino acid sequences
of ketosynthase domains prom various type I PKS were optimized
for myxobacterial DNA by comparison to a known myxobacterial
biosynthetic gene cluster (Schupp et al., J. Bacteriol. 177,
3673-3679 [1995]) resulting in primer
KSlUp (5'-
C/A) GIGA (A/G) GCI (A/C/T) (A/T) I (C/G) (C/A) IATGGA (C/T) CCICA (A/G) CAI
(A/C) G-3 ' ) and
KSL1 . (5' -GG {A/G) TCICCIA (A/G) I (G/C) (T/A) IGTICCIGTICC {A/G) TG-3' ) .
PCR-primers TGD (5'-
T (A/T) (C/T) CGIACIGGIGA (C/T) (C/T) (G/T) IG (G/T) ICG-3' ) and
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
23
LGG (5'-
A (A/T) IGA (A/G) (G/T) (G/C) ICCICCI (A/G) (A/G) (G/C) I (A/C) (A/G) AA (A/G
)AA-3')
directed to genes encoding adenylation modules have been de-
scribed by Turgay et al. (Pept. Res. 7, 238-241 [1994]). PCR
reaction mixtures with a final volume of 25 ~l contained 0.1
template DNA, 0.2 U Taq DNA-polymerase (Gibco BRL, Eggenstein,
Germany), 5 ~mol dNTP, 5% dimethyl sulfoxide (Sigma), 1.5 mM
MgCl2, 25 pmol of each primer and the appropriate reaction
.0 buffer supplied by Gibco BRL. Chromosomal DNA of S. cellulosum
was used as template,. Additionally, chromosomal DNA of Myxococ-
cus fulvus was used with PS primers. Reactions were carried out
in an Eppendorf Mastercyler Gradient (Eppendorf, Germany) using
the following conditions: denaturation 30 s at 97°C, annealing
30 s at 55°C, extension 60 s at 72°C for a total of 30 cycles.
The formation of ca. 700 by fragments using the KS primers and
of ca. 350 by fragments with the PS primers were confirmed by
0.8% agarose gel electrophoresis. Fragments of independent PCR
reactions were ligated into vector pCR2.1TOP0 using the TOPO TA
Cloning kit (Invitrogen, Leek, The Netherlands) according to
the manufacturer's protocol and transformed into E. coli XL1-
Blue. Sequencing of the resulting plasmids and analysis of the
deduced amino acid sequence revealed three different KS frag-
ments, designated pM008.4, pM008.6, pM008.7, one PS fragment
(pAPsl) corresponding to S. cellulosum and one PS fragment
(pDPsl) obtained with chromosomal DNA of M. fulvus. The PCR
fragments were re-isolated by digestion wish EcoRI from the
plasmids pM008.4, pM008.6, and pM008.7, labeled, pooled and
used as gene probes in hybridization experiments as described
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
24
below. The same procedure was performed with the PS fragments
of pAPsl and pDPsl.
Hybridization with PKS and PS specific DNA probes (see
above) was carried out using the DIG nonradioactive labeling
and detection kit (Boehringer Mannheim, Germany) and performed
according to the supplier's manual using buffer containing 50%
formamide. The membranes were hybridized in plastic bags con-
taining approx. 10 ml of hybridization solution at 39°C over-
night. Unspecific binding of probes was removed by 2 wash steps
with 2 x SSC, 0.1% SDS at room temperature for 20 min. and one
stringent wash step with 0.5 x SSC, 0.1% SDS at 60°C for 20
min. Detection of hybridizing DNA fragments was performed with
the above mentioned system according to the manufacturer's pro-
tocol using CSPD as chemiluminescent substrate. The signals
were recorded by exposure of the treated membrane to Hyperfilm
ECL (Amersham Life Science, Little Chalfont, England) which was
developed in appropriate time intervals.
71 signals were detected with the PKS specific gene probe.
On the duplicate filters 35 signals were obtained with the PS
specific gene probe of which 7 were already known from the PKS
hybridization experiment. These recombinant cosmids harbored
PKS- and PS-encoding genes. In order to corroborate these re-
sults PCR experiments were performed with DNA of the 7 recombi-
nant cosmids as template and PKS (KSlUp, KSD1) and PS specific
primers (TGD, LGG) generating fragments of the expected size of
approx. 700 by and 350 bp, respectively (primers and reaction
conditions see above).
A comparison of the restriction fragment patterns of the
DNA from the 7 recombinant cosmids carrying PKS and PS genes
digested by BamHI facili~ated an arrangement of the cosmids in
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
3 groups. They were rep=esented by cosmids designated A2 and
A5. The remaining group was represented by pEPOcos6. Therefore,
A2 and A5 represented gccd candidates for further DNA sequence
analysis because they carry both PKS and PS genes.
5
D. Random "shotgun" sequencing of recombinant cosmids and plas-
mids
1. Library construction
10 a. pEPOcos6, pEPOcos8, A5, and Sau4: pEPOcos6 and pEPOcos7
were sequenced to comple=ion, and contiguous sequence data and
analysis for these over~apping cosmids is presented below for
the "cosh region" (cf. M aims 7 and 9). Sequencing of cosmid
A5, pEPOcos8 and plasmid Sau4 was taken to the point of large
15 contiguous sequences (ccntigs) representing the S. cellulosum
insert; sequence and analysis presented below (cf. claims 10 to
15) .
Randomly sheared l;braries were constructed for cosmids
and plasmids of interest using a protocol similar to that of of
20 Fleischmann et al., 19~~ (Science 269, 496) and modified in
Fraser et al., 1995 (Sc=ence 370, 397). Briefly, Qiagen-column
purified cosmid DNA (-1G fig) was sheared to a size of approxi-
mately 2 kb and the DNA end-repaired using BAL31 nuclease. The
DNA was gel-purified after electrophoresis through a 0.75% low-
25 melting temperature agarose gel containing 0.5 ~g/ml ethidium
bromide in lX TAE buffer run at 80 V for 2 hours. The volume of
the low-melt agarose gem slice was estimated by adding the gel
slice to a microfuge tine and weighing, then 0.1 vol. of 3 M
sodium acetate (pH 7) ~:s added and the agarose incubated at
60°C. Tne temperature :~as equilibrated to 37°C, and DNA ex-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
26
tracted twice using an equal volume of buffered phenol (Life
Technologies). The aqueous phase was transferred and extracted
once with an equal volume of chloroform, then ethanol preci-
pated by the addition of 2 vol. cold 100% ethanol. DNA was con-
s centrated by spinning at 16,000 x g in a microcentrifuge. The
DNA pellet was washed with 1 ml 70% ethanol and resuspended in
100 ~,l of O.1X TE. The DNA was ligated to SmaI-digested, phos-
phatase-treated pUCl8 vector (Pharmacia), and single insert re-
combinants isolated by gel-purification of the band containing
vector plus a single insert, followed by T4 polymerase polish-
inc, and a final intramolecular ligation of the vector-plus-
single-insert DNA. This final ligation represents a library of
highly random ca. 2 kb fragments that was used for shotgun se-
quencing of the ca. 40 kb cosmids or ca. 10 kb plasmids.
b. Cosmid A2: Cosmid DNA with inserts of S. cellulosum was
isolated by an alkaline lysis procedure and purified with Ma-
cherey Nagel columns (Machery and Nagel GmbH and CoKG, Diiren,
Germany) using manufacturer's recommendation. Purified Cosmid
DNA was sonicated, end-repaired using T4 DNA Polymerase (Boe-
hringer Mannheim, Germany). After gel-purification fragments of
a size of approximately 2 kb were ligated into SmaI-digested,
phcsphatase-treated pTZl8R vector (Pharmacia). The ligation
represents a library of highly random ca. 2 kb fragments that
was used for shotgun sequencing of the ca. 40 kb cosmid.
2. Sequencing and assembly
a. pEPOcos6, pEPOcos8, Sau4, and A5: DNA (1 ~,1 of 100 ~.l
total in the library) was transformed into E, coli by electro
po=ation (20 ~1 of Electromax DH10B cells from Life Technolo
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
27
gies) and cells spread onto LB plates containing 50 ~.g/ml ampi-
cillin. After growth overnight at 37 °C, transformants (ca. 300-
3000 CFU total) were tranfered to 96-well growth blocks and
shaken overnight at 37°C in 1.3 ml LB medium with 50 ~.g/ml am-
picillin. Templates were prepared from these cells by an alka-
line lysis procedure (Qiagen QiaQuick Turbo Prep) to yield pu-
rified, double-stranded plasmid DNA. Cycle-sequencing of the
plasmid templates was performed using universal forward and re-
verse primers and BigDye Terminator sequencing kits (Applied
.0 Biosystems), using the manufacturer's recommendations, then re-
solved using an ABI377 automated sequencer. Sequences were ed-
ited using Phred, then assembled into larger contiguous se-
quences using Phrap (Phil Green, University of Washington, St.
Louis, MO).
b. Cosmid A2: DNA (1 ~1 of 20 ~tl total in the ligation)
was transformed into E. coli DH10B by electroporation and cells
were spread onto LB agar medium containing 50 mg/ml ampicillin.
After growth for 18 hr at 37°C, transformants were transferred
to 96-well growth blocks and shaken overnight at 37°C in 1.3 ml
2x YT medium with 50 mg/ml ampicillin. Templates were prepared
from these cells by an alkaline lysis procedure (Qiagen Qia-
quick Turbo Prep) to yield purified, double-stranded plasmid
DNA. Cycle-sequencing of the plasmid templates was performed
using universal forward and reverse primers and Big Dye Termi-
nator sequencing kits (PEBiosystems) or Thermo Sequenase fluo-
rescent labelled primer cycle sequencing kit (Amersham Pharma-
cia Biotech) using the manufacturer's protocols. In the shotgun
phase of a cosmid, identical amounts of samples were sequenced
either by dye-primer or dye-terminator chemistries (Pharmacia,
PE Biosystems). Data were collected using Licor and ABI 377
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
28
automated sequencers and assembled with the GAP4 program (Bon-
field, Smith, Staden, Nucl. Acids Res. 23, 4992-4999 [1995]).
Gaps were closed using custom made primers (MWG-Biotech) on
plasmid templates or PCR products in combination with dye-
terminators.
E. Bioinformatic Methods
1. Open reading frame (ORF) identification
ORFs were identified ==. the pEPOcos6 region using the OMIGA
1.1.2 (GCG 0.4D) program ~rom Oxford Molecular Limited. Default
values were used (Stanaard genetic code, all ORFs over 50
bases) to generate ORFs; analysis of these results lead to the
list of 14 highest quality ORFs as defined in claim 9. Other
ORFs, genes, or genetic elements may be found in the pEPOcos6
insert that have not yep been annotated. In addition to hand-
editing of the OMIGA-ger_erated data, the MAGPIE automated ge-
nome analysis tool:
(http://genomes.rockefel=.er.edu/maqpie/maqpie.html)
was used to identify genes for all the sequenced cosmids and
plasmids. ORFs identified in this manner are presented as both
nucleotide and peptide files below.
For cosmids A2 and A5, ORFs have been identified within
the DNA sequences of A5 (contigs 10, 11, 12) and of A2 using
the FramePlot analysis program from Ishikawa and Hotta (FEMS
Microbiol. Lett., 174, 251-253 [1999] public available under
[http://www.nih.go.jp/~j~;n/cgi-bin/frameplot.pl] which is based
on positional base preference in codons typical for organisms
having genomes with a hiJh G + C content (Bibb et al., Gene 30,
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
29
157-166 [1984]). Default parameters using ATG and GTG as start
codons were used. The deduced amino acid sequence of predicted
ORFs were compared with protein databases (GenBank, CDS trans-
lations, PDB, SwissProt, PIR, PRF) using BLASTP (Altschul et
al., Nucleic Acids Res., 25, 3389-3402 [1997]). Additionally,
high scoring amino acid sequences were analyzed using the Pfam
program [http://www.sanger.ac.uk/Software/Pfam/], which identi-
fied specific domain ~~ructures of the submitted proteins
(Bateman et al. Nucleic __~ids Res., 27, 260-262 [1999]).
2. BLAST searches
BLASTP2 similarity see-ches were performed using the peptide
files from the above OR= identification strategy as query se-
quences. Searches were rerformed using the in-house Bioinfor-
matics BLASTP2 (Version: 3LASTP 2.Oa19MP-WashU) web page at the
Bristol-Myers Squibb Pha=maceutical Research Institute (allows
BlastN2, BlastP2, BlastX~, TblastN, and TBlastX searches). In
addition, peptide files generated by the MAGPIE analysis were
automatically searched using a FASTA algorithm.
3. Best match and probable identification
A:~alysis of the BLAS==2 and FASTA output led to an assign-
mer.~ of a best match any probable function. The best match was
usually the top scoring -.-.atch, although sometimes another match
was given because it was a more relevant homolog, or no match
was found with a signif~.vance greater than >e-4. Probable func-
tic~ represents the bes~ estimate of function given the initial
ana~ysis of the BLAST da=a and the published literature regard-
ing the best match, ana -:gay not necessarily represent the true
fur_ction of the gene product (hypothetical proteins are of un-
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
known function). A higher probability score indicates a higher
liklihood that the probable function corresponds to that of the
best match; e.g., the polyketide synthase matches are all above
e-100, and given the very high significance scores are presumed
5 to function as polyketide synthases (as are the high scoring
peptide synthetases).
The following is a summary of the sequence data from the
pEPOcos6 region, pEPOcosB, A5, Sau4 and A2.
10 a. Data from pEPOcos6 region:
Summary: A large PKS/PS cluster spanning multiple cosmids.
An IS element (designated IS-Scl here) is found in the cluster
- this may be a potential tool for genetic analysis of Soran-
15 gium.
Statistics: Sequence was assembled from over 2000 random
sequences (forward and reverse reads of the ca. 2 kb cloned
fragments derived).
47,713 nucleotides of contiguous sequence (no pFD666 vec-
tor included)
DNA sequence data are as defined in claim 7.
Note: pEPOcos6 ORF7 sequences (cf. claim 9): the predicted
N-terminus of ORF7 shows 145 nucleotide overlap with ORF6.
Note: pEPOcos6 ORF8 sequences (cf. claim 9): >pEPO-
cosh ORF8.seq ("ORF9 up" in Fig.2)
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
31
67.3% G+C
Table 3 shows ORF data summary. Note: pEPOcos6-ORFi.seq is
truncated at its 5' end; correspondingly pEPOcos6 ORFi.pep is
truncated at its N-terminus.
b. Data from pEPOcos8 region:
Summary: Two PKS genes found on a cosmid. A Tn1000 inser-
tion is also found (occ::rred during E. coli propagation). No
peptide synthetase genes were found; one P450 hydroxylase was
identified.
Statistics: 1952 random sequence reads from the pEPOcos8
library were assembled using phrap, with 7.024 of the sequences
assembling into 57 contigs. 12 of these contigs were chosen
(totaling 56,537 bp) which each contained >6 reads and con-
sisted of about 1000 by or more. The sequences of these 12
contigs and the associated ORFs are given below.
DNA sequence data from contigs are as definded in claim
l0. Table 4 shows more data.
pEPOcos8 protein da~a are as defined :in claim 11, i.e. for
selected ORFs (polyketide synthase, peptide synthetases, or
ORFs with high similarity to known genes).
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
32
c. Data from cosmid A5 insert:
Summary: A cluster ~= PKS and PS genes found on the cos-
mid. Other genes possibl-.- involved in this secondary metabolite
production include a dow.~stream lipoxygenase gene higly similar
to eukaryotic orthologs.
Statistics: 880 ranaom sequence reads from the A5 library
were assembled using phrap, with 530 of the sequences assem-
bling into 12 contigs. _ ~,f these contigs were chosen (totaling
41,556 bp) which each ~cntained >100 reads and consisted of
about 9000 by or more. T:~~ sequences of these 3 contigs and the
associated ORFs are gives below.
DNA sequence data f=~m contigs are as defined in claim 12.
Table 5 shows more data.
Protein sequence da~a from selected A5 ORFs are as defined
in claim 13.
d. Data from plasmid Sau4 insert:
Summary: Insert cor_~ains PKS genes on two large contigs -
most similar to the sora~~en PKS gene from Sorangium.
Statistics: 565 random sequence reads from the Sau4 li-
brary were assembled us~.ng phrap, with 84 of the sequences as-
sembling into 18 contigs . 2 of these contigs were chosen (to-
taling 6596 bp) which eac:~ contained >10 reads and consisted of
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
33
about 1000 by or more. The sequences of these 2 contigs and
the associated ORFs are given below.
DNA sequence data from plasmid Sau4 contigs are as defined
in claim 14. Table 6 shows more data.
Protein sequence data from selected plasmid Sau4 ORFs are
as defined in claim 15.
e. Data from cosmid A2
Table 7 shows ORF data summary
F. Construction of suitable recombinant expression vectors
1. Expression in Myxobacteria
Heterologous expression of the ORFs shown in Figure 1 is
performed by using a derivative of plasmid pSUP102 (Simon, R.,
Priefer, U., Pizhler, A., Methods in Enzymology (1986), vol.
118, pp. 643-659). In this plasmid the gene for chloramphenicol
resistance is changed for a cassette comprising the gene for
streptomycin resistance and the promoter element of the Tn5
transposon. Short homologous genomic DNA segments from the host
organism are ligated with the DNA sequences of Figure 1 and
wit:? efficient regulatory elements into, for example, the EcoRI
restriction site of the vector. Following amplifiction of the
vectors in Escherichia coli the DNA is transfered by electropo
ration of the host cells or by conjugation with Escherichia
cola S17-I (Simon, R., Priefer, U., Puhler, A., Biotechnology
(1983), vol. l, pp. 784-791).
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
34
By means of the tetracycline or streptomycin resistance,
respectively, mediated by the vector the host cells are checked
for integration of recombinant plasmid DNA into the chromosome
by homologous recombination.
2. Expression in Streptomyces cells
Heterologous expression of the ORFs shown in Figure 1 is
performed by using bifunctional Strepomyces-Escherichia coli
cosmids pKU206 and pOJ466.
3. Expression in Escherichia coli cells
Heterologous expression of the ORFs shown in Figure 1 is
performed by using "bacterial artificial chromosomes", cosmids
(for example Supercos, Stratagene GmbH, Heidelberg) and T7 ex-
pression systems (Stratagene GmbH, Heidelberg; New England Bio-
labs Schwalbach, FRG). Expression of recombinant enzymes occurs
in Escherichia coli cells constitutively expressing phosphopan-
tetheinyl transferase required for the formation of holoenzyme
polyketide synthetases and polypeptide synthetases.
CA 02346499 2001-04-09
WO OO/Z2139 PCTNS99I23535
N
0
,J
U
N
N
ro
C
.,.I E
x
o
G U
v
z a~ ,J
o
H ~~ a) ..
ro U O ~ C -CC .. a)N N a)a) C G
C a~C v -.I ..-a-~ a~ -I1.1 G
H v ro In~na~ a)a~a7 v~cn m m m a~o,
L-~Zn.t-~y~ ro~pa~ W L ~ roroU: rororo 1-iI '
f~ O rotnO .C.CO V O O .c ~C .C.C.c O N
H Sr 1~-.-i ~
a ~ a a a a a ~ ~ .N+~a~ a C n
H ~ a.~nn. c a a c,a. c c a~ c ..c o. G>
z ~, x a~ ~,~, ~,~,s :~,:>, o
w ,Js-~~ m u~-r r,r;,~ ~n~n~ ~n~nm rl
C roQ)
a ro -~-4iU C:U U a)a)7. N N a) U y r.
w e~~ ..~ .a..-a'O-Ocr,'z~'D'D w
o .~ .....~L -.~:.~~ ..~-., .~-.~.~,~ ~ 'J
a o a~ L :JN Q1C~C7 :1J G! 11~ 11 N
;. m a;~ a;v m .crJ I
~, s...:L _1'.111 O O .~ ~C.Y L G r
~ O :r.7,G O :>,?,.- ~,:u,>, O --a
LL J ~ G. C. .~..: 7, "~-' p O O
ro -,.~.~,Q G.~ J-.. ~ ... ~ ~ ~ L
'a.~ .G w .C _-.:.1~..w ' w
ro ., G.
O C
O
U
O
_ _ ..-, O U
---. O~ O r-i -,,--,'--~ O ,-,r..-..-. a.
Q .-. M c W D O O~
' N N 01r-~ ~ ~ '-IN ~ ~ M ~ M
V N d.1 I ~
I ~ ~ I ~ ~ I I I I .-'l.7N e-i I '[j O
O ~ ._, ,...J Q7N U~ N OJ.--iH N ,~ 07
I ,-.I I I I _ N U
U ~ .~ ~ ~ .C ~ ~ a N N ~ FC
p cn v cn ~ ~ c
E U cnG.U v~~n cnr-c~ .-,O rl U >~ ro
U O N U .7. H H H U O X I
H rs C'J tW C J.~~ ~ ~ ~ N u~N ro~ U 'O
V m ao~nro cnr ro c~rx~l ~~inr .io ,-~ i
f ~ ~ E I ~ E I I _,- E s, ro
.. ~ ~ a,O x o~ G.O C7 < a~v,o~ ~
D O r O tnr'7O ~ ~ ~ a C o O O O Z :.~ I
W ~~ fsuoC ~Gm C c a,cr :G.-,iv Cs.GJG, C O
L4 ~C ~tO -- c1c~-- ? >.r G.f-..~
C
ro
:~
N .~ r...y:..-,~ :O'(w~o
Gl W f~N O',N iI7~ O N C' i H
r-i.-~G,.-~a7v,O N O IW n ~ ~O 0.-ir-fV7 ...47
~p:v;.-,G .-~N a C)
J~ tW O Q~v, f''1O NO M M ~ o~01r W r .-~I; :-'~ y,
N ~f1f1N 0007CJ l0r-t.-aN vl lulDf~ ~ tf1 ,J
N N H N ~
p C~ W
O
W.
Id ~ u7N M O~N O f~OJM (''701.-aa,~f1c''1O .-i G
rl~Il.-ir-1C'~ ~ -1O N ("1C7 C'~ ~ ~TN rJ
GI N tf1f!1N 1flt.~r tf1.~,~ ~--n1f~..~7.~ v,OO H O
N N ~ N r, 4.
tn (~ s .. M _
u1O~G1 10cf,.-~c,~ .-I M N U1Q1 G' a' J
N C' , lD err.
p ( tf7O l0 N r ~ f ~ O .-~r, f!1N N N
I~ CO N l0M COO'~ ~.Dd'W c u7 i_-.
rl y -1r1 ('~N N r-1 W ': c f-~f1 r-~
:J
d ~ _ u1~ O .-IN '~ lDa _ G c ~ N O'~x (~
EI ~ .~ U1
f t~Inu-7O N u~ O~tf1C M ~ C~N u~ -,I~ ro
N M N t~O .-it1~' .-,.-~~ tDC~:CJ c 01u'1O v7 Gi '-'
N (hv, N C,l0 v,1DOr c~-u'1. N d1~f'1~
p ~ .--''-I,-1,--~,-,.-,N t~::~r'7v, crc C.
r1 .-~H CV ~ f W l0~ ~ O b t17i~~ r-1Q1to .~. L
AI f~-1O .J ~a,01 l0O ri ('~N r" 1D01('~101r-i v
W r r'1V, ('JO H W Lf1pp W lDN N ~ O u'1O ~ .
(l, N fl v :J~ u-ltO~p ~OI W: ~ N O ~h~O G.
n .-~Irir-r,~ r-i~ N N c'~~, Q,c .-
r ~-f
-72' M
N M
C H N M M a'
r-1 ~ .-~N t".tQ, 1f1lDr~ ~ .~~..-CDG'~.-...--~.~.-,H .~ 17
LL LLLLIL ILIL:L W ' I--.rW ~ W W LLL.LW It.
H C7 O O O O O O O O lLC C O O O O O O O
O
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO OOI22139 PCTNS99/23535
36
x
.-i
a.
r, v
rn
r
N
O U
ro
c ~ c w
N
QI w 1
tn '~'E
N O m
a.
a ~ : '
_ . U
C ro
~ o
.,~ ,~ a o
b a~
a~ > a,
o C
a ~ o
.~ a~
a v
x
o w .-'
a o
v r, ,--~.-,
o a
p a ro
j ,n ~ O
ro
a +~
m ~ ~ ro ro
r-,a~
c 3 ~ ...aa
o c
o ~, ro
U
G
.. .>:~ N
L ro
w C C
C O
..-1 1 L rl
N
1a ~ L 1~
ro ro
E. " ~ .-,
. .~ ~ c1 ,-.1
a~
s-
0
-o '' _'''v Q
~ L
U "' u~
N CJ c
C :i CL
~~ ~ a
,.
" -r -~1a.
C
C sa O '-i
C r
N C v ~, w r
~ ~ N
Q iJ O 1a
.,
1
i-~ . ~
.
~9 N ~ C
.p E
G ~ N
C C a r-i
.i!t)~ 9
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO 00/22139 PCTNS99/23535
37
0
U
a
N
.O
G
C
O ...
-.-1
i
W O
N O
U ~ O N
.1 b
O 1~
.-I JJC C
1~ G
.~ ?i:.
Q
G~
~ .U Ti
1 ..a ..~1
~'C
~i N X
i ?,2
I -I O
'Q O m
O
i-1
r
N
+ i
O p O fv
Lt O O ..r
O r~
U 1 ~ N
t11 N CJ ''i
.-,i~.-r u)
X
C Q.
U' W
y~ U N Ol
O b ~CO
~ u7~. 00
p. .-a
W LJI N - N
U71 Y~-- O
.L~ ~ ...
O ~ O N 01
~y a, r ,n
b OD. ~ N
~I N N
b
C QJ M a~O
~ ~; r
v
o ~ r
y 00N ~
~1
. tnc~ O
1
('n r C ~ O
l047 N O
G i m ".,.-ir
O
U
I
N O I c N N
N v7 M ~ N
W
W r ~o w o~r c~r~,-,r ,-,u~ v N i
N ~ .--tv r l0r ~ N r ~f7N v U7 ~ i
~l O N O~Q~N r'1ODCO N O Q~ .-~ tn .-iN
fll W ,-~~ .-iN N ~ T r tf1O '~ ~' I
(C N '~ "'-'~
H b O C~ O O
N t11 U ~ O O O O
r~ .a ooavO ~ N r~ ~rm o r y~ I I I
O Y~ ~o
U ~' V' S crtf11f7U1Lfl~ lf~1n U1 U vo r r
Q a ~ i 11'7~'7U1
p, O~ ~ CPtnrT CTU~CT CTO~Cs ~ ~"'i '-a
W _~ _~ ~ ,~_,..r.~..1.~ .~.,.-a.1 d N .'..i .'-r
tW 1JLJ J.-)AJl~ .J-1i J-11J N I 1.1 ~JJ-~
G C ~ G C C C C C ' C C C '. ~ G
b tl1 O O O O O O O O O G O O ~ O O O
~ U U U U U U U U U O V U .~1 41 V U V
V
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO OO/Z2139 PCT/US99/23535
38
c
~,
x
0
o
.r.,
o r,
a
w w
0
0
0
U C
O _y
L
U
0
(V
<T
N
W
U o1
O
r
N
x
w
U
01 O~ t!1 N n-1 C' OJ ~ v' r N O ~' N dD ~T C' ~
C ri C' N .-i r OD R'~ (''1 N 01 r l'~ O r tD M C~ N
e-1 cr N N C M N rt r-1 M amfl N M N '-I r-i M N
0 o m a~ ~ ~ r oo m c a~ M ~n r r mo o~ o
u»o ~n ~a M N ~o ~ o m r ~ o N M o~ o
c N r ~o N ,~ co c ~r c~ c r r Q' ~ °' °' ° ~o
00 .-~ v m co r m m c o m o
m u~ r o r a m ~ a~ o cv .-~ ~ M M o~ 4~ ~ o
r r M w ~ o mo 0 0, o, o r .-, ~~ M c r
O~ M M 0 ~O ~9 r o .-~ O N N M M M C' a' ~
~D O 00 Q1 01 OD Q1 ~ N r-~ ri f-i .--1 r-i ~ '-1 ri .-i r-1
V' lD c N N c v-i u'~ r r O O r'7 Q' r-1 O ~ 00
N ~ r ~ M N N c0 OD CO N M O1 W r O M r
c .-a O '~' c~ r ~ O lD 00 W7 r O O M 01 O 10 N
r r av w o~ o~ o ~ o r, .-~ M M ~r c M c c u~
~ r o ~ N M r o~ o .-~ M °' r m °' '-~ N M '°
N N M M M M M M v~ c' c
O O O O O O O O O O O O O O O O O O O
r r r r r r r r r r r r r r r r r r r
m umn um ~n u~ um mmn m ~n u~ umn u~
CT CT Q' IT O~ CT CT CT ~ Cr CT CSC Cr ~ CT ~T tr~ ~ O~
.,~ ..-,
L J..J L 1J l~ J-~ Y 1~ l~ 1= L J..~ 1.J J.J 1J 1J .4.1 1~ 1J
C C C C C C C C C C C C C " C C C C
O O O O O O O O O O O O O O O C O O O
U V U U U U U V U U U U U U U U U U V
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO 00!22139 PCT/US99/23535
39
Table: 5. A5 assembly analysis summary (continued)
a. pEPOcosB assemblies
~ ~ a
E m a
'
e9~' v v v ~ c
ov Y a Q m
d ~~ ?
~~c~cc
~' in :n Q :o
~ ~ -v
v N
m ~ of
~
v!a c v ~ ~ r> '
'o
a , v ~ 6o r am
o ~ ~ a ~
3 c
a
(/lV N ~ i r- ~ t~ r
I r O ~ Q1
C Q L1 of c
~n Z '; c~
Y N (ti
~
O ~ ~
_
O
~ C l9 m h
= ~ ~ rVNC
~
( O r ~t0~ N fht fp
~ f0 N O
O ~ f9 00
~ In
~
t0 t0 r ~ c0 ~ ef M N
Vi N C ~ O a
~ ~
~ N
Ln O1 ~ C O~ e
N -
'CC c ~ ~ N ;o p ~ v
E n ~ Q
~
C C ~
'
4=t~ fl :~ : ~~ C p O c0
~ 07 V G1 ' U ~ O M
U ~
r ~ ~ C U O C C
Q C
~
lL ~ .D C d O 'D O
~ C O ~
N
~ ~ I ~ D
g
N
a'
Q Q E ~ . Q a
c D v Q o :o
D $ ~ a
N N N dl
N N O In VIO
N
Ul _ N f ~ ~~V N
M ~ L b
NN LO
_ d
C N N N N ~ C N C ~a tG O
. d C U ~ ~
C _
~ ~
>
~
r Y i -. -. T . y~ C0 G (0
j, t :~ N . C 41
~ .' ~
C C U
C ?.
of
L
C O C Tll7
j ~ N NE O
a Y
N d y fn Vl ~ ~ t E C Of
f' N o - j, d du. o v
= ~
-
~
d E c d d g a :"-' ~
d :o
v_ v Q Y ~ a -o a
a ~ a
~ ~
~
a n. a >.>..c a
a
a a o
a a
a n..
O V t1~~ O ~ ~ O)
~
r r r ~ O
r N
t0 L N O _ N O n ~ ~ ~ ~ N
N ~ ~
O 01
GO ' f0( c~v h v
D Nri~
t
rr V r ~ N _ V M O U7 _ r tt r
f .C r O O h al U w U U
L U r
Vl ~ ~ V O ~ O r O V O N
O .U..~ p7 O
GO p f0 fU f0
CD (D f9
E '~ m ~' o U M E E E E
''' E m E
U ~ ~ ~ m d
.,
c' M Q tn U Q dZ c~cccU
'7 p Q ~ U
co. a.cUU
.. _.
0 0 0 a s ~ ~ ~= 0 0 0 0
0 o a 0 0
0 '
0
'h V)t0 O O r a0 ~ CO c0 ~ W N .- aD
w r N ~ n W c
7
. O I' ~ u7 O h.
O O N N f'1
O c'~1
O O O O Q7
0 0
CO CO Q7 07 07 ~
O O f37
tO ,~ tn N ~ h
O r
Q1 ~ p
~ O
r
A
N O r 00 O ~I7 07 (O N M
tn r N 00 O
tn C
O
N O f1 _ h 1n 01 '~ t' ~ O
00 Q tn O G~ 00 N
7 In O GO Q7
~ _ _ h c'~ Cr7(D d ~ T h h h r
N h N N r r
'V ~
r m v co o h c~ co c v
o v co o~ o ~ m
_ N OO G7 01 h
tC~O ~ CO
~ O O O)
cD
W tn h tn N N O
tf ~ tn N tn h OD
V'
N M h N tn (Ja0
fD O c~
!- N V r M ~ lO CD r N V ~ f0
C~ 117N f7 h W 07
O
tp
f"? ~ tA
a ~ o
0 0
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO OO/Z2139 PCT/US99/23535
ro
L
Q7
N
ro
C
C tn
N
b
a)
~ ro N.
v
o ~' n.
a>
r,
0
n.
a~ i
u1
u1
" C
i
U
.-, C1
C _
O
U M
-i C
.-a
ro 1 ~1
Q7
w
O O
~~y~ N
U1
M a'
Q, O O
t0 01f d' .y.N ~ ~ 01 M .--i171e-O~Q1 f~.--mT (~.-a~~ a~
t0 Ir1v'N M W O !'~O M r i.nO v~O .-I0107 O tDM c
~ f~.--1M M M OD C'M N .-~N N N u1N .-W1~ ~ M a'
O ~ 01CO c CV l0If1c'1D~ tf7
L~ 00c>W O W O M lG~O N l0O if1~'C <t'1~t17N m O M
N c f~ O O v ~rO N c ~ m .-av0M u~1~~ ~ O M l0
x N V'Q1 .--1r~N ~ 01f~ M W f~ 10r11D M r1~ rlr-1ri
b . M N h 01~ O OD01 Lf1!~~O 01tf1~O t~ I~~ a' 01
M N N O ~ ~ O7 O M r-r01O1c' N M O f ~ ~ ~ O '"1
COM 01 Q1N t!'7tD.-aN N f~O O ~ CO r-I111(~ C1i!1N 1f~
111M N M M lC u7~Dt~ f~t~ a161CO Q' N ~ N M M
U
.-1O l0 M r c~ M M O c N 01 tl'1~'II1 t0M O l0f~ M
l007!~ tf~W CO M c O tf7m .-1,-;01M O N M O O7O r-1
~ aom o0 01M .-,o o m a101a ~rr ~ r Lr10 ~wfmn N
N N M N c w f~. ~O l0v7 f~('~61 f'~N ,-aM M N
.-It0f~ ODQ1M 1:Jf CL701O .-1M s'f'-r-aN ~1'U'1r O7 r-t
O O O O O .--~r~.--W"~N N N N N O O O O O O .-i
O O O O O O O O O O O O O O O O O O O O O O
~ I ~ ~ ~ , ~ i i ~ ~ 1 ~I~I ~I~I~I ~i
IC) e-1r-1rl v-1,--I~ r1v-1r1 ri.-1n-l'-1e-1rW-iri r-if-1rl N
G) Q' ~
.~1.-1.-1w-~i.-1w4 rl.i.-I rl.-I.1.1ri .~wl-n n .1wl ri
d
.1 N a) lJ.u.u ~ :.~i~ .ua m -~~ ~ ~ .ua.~:~.u:~ 1J~ L a~
C C C C C C C C C C C C r' C C C C C '" C ~ C C
H ,L~ O U V U U U V U C.~U C'U U U U V U U W % U U U
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO 0022139 PCT/US99/23535
41
w
z
w
v v v
x
ro roro ro
a
" cu
C C v .~ 1
~ a
m N ~r C w
C ~,,
v w ~ ~' ~
.o .a v z
"' '.~v -a o
.d .,~ o
N ~ .ri ~ H
X y L ~ x
>, ~,a v
0 0 ~' '
a
N
01O N
N O U
.-1M
C I I E
-..1 v v O
V ~ ~ L
>~ U C N
ro a .a a
w O U
-..a 17?~ cO
_
la C ~ V c
a~O
N C M
I
ri ~O
1 N I 'J
W ? W 1 W
N
N Q. O O
O,OD
o N O 0
r m
M tf7r N
N ODV' W x
N M O O
h a~I~. w ~l
r~ N fw ~
M O~
N M c .--~ O 07v~ l0O W v v'N O t'~N r-r(V l0COCO Q1N 1nO C'
m M 01N O ~ O ~ u7V~ra rrO aD 0701O~ !~O O~OWI? I~v N M c
CO~ M r-rM N ~ ~ c tf1M ~rr-W ~ ~ ~ .-~N M M v ~ N N N tn
1
01N ~ N O U7 00M tWO 01 .-W~ f
v~O W c c M f~~T ~OU'1t'~M tnV' M V'01 lDCT O~01f O Q1O~M M
l0M r-1~DN M N u',M t0~ M ~-1!~~c O~(~ r-aO ~ .--~M v'N h ~ l0
N .-iH ('~01 1DM n-ir-1r-~O~ r-1M r-it17tWf1 tn~D r-1,-ir-rU7f~lDlD H
tWM
.--~O ~ 00O l0N u1 ~ H 01lDr--~~1.-~O ~1M l061~-,O N C'WD O
,~.-1M In01 M f~C' ~iLf7l0 V~N r-1lDM N V'(~-O~61M r-1N rl tn
CDN f'-N O N f 1!7 l0f'Wt7O~01 O N ~ inM O CO~''1O OJM fW fi
M In lDl0OD .-~.-~ri raH .--~c N tl7u'7W 1f'7tD r W f C~I~. I~ O~
01 M f~
O~ri .-i01N M 61 N f~ f~'1r O~ M N (\~1D~Il10~ ~'7O~r';f~N 1D
.-1tf101M f~M r-i~.OM H ODf'-tf7N M cW f'1~,OO~O1tI1c ~ 01O H
C U'1~f'7OO(~ t0f~ M M 00f~19 ~f?l0Ol O f'~ODO 01 ~ O Q1V~ Q~
l0l0 Lnal.-~,-1.-a '-1 N N riT C'C'C' 1D~ tnf'~v'1, f~. OD f'-
N l0 Q7.-WD ODQ~.-iN M u~ ~Of~-Ov .-~N lh ~ru) lDf~O~ O .-.N f''1v
.-1.-Wr N N v~v O O O O O O O ~ ~ ~ ~ .-~~ ~ ~ N N N N N
O O O O O O O O O O O O O O O O O O O O O O O O,CIOI Oi
~ i
~ .-1.-~t~-i.-I.aN N N calN N N N N N N N N N N N N N N N
.-W--W--1rir1 r-1e--ft-1.-~r-ar1 r-W-1r~ '-I.-1v-Ir-Ir-1rir-~H rln r-ir-1r-i
CTO~ CT>Tp~ CTtr~:T ~ O~tn O~CrU~ ~ is~ rTCT CT~ O~ CT~ ~ O~ aT
.,~.,1.,i.,.~.,1.~.,1.,1.,.1.,~.~ .,-
~.,.~.,~.,a.,1.,~.,~.,~.~.~...~.,~.,1.,.~.~ .,1
111-11J1JJ-1.1~1JL 1-~1~1-1J.J1.~1J L tJ~1 J.J1~ L .!J.J JJL 1JL 1J
C C C C C C C C C C C C ~ C C " C C C C C ~ C C -'C C
O O O O O O O O O O O O O O O O O O O O O O O O O O O
U U U U U U U U U U V U V U U V CJ U U U U V U U U U U
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO OOIIZ139 PCTNS99/23535
42
o~u~~r o c ~
M v~~ ~ r
M a V' M l0t0
O OD~f1 O .-a
N M c M N u'1
p M N M O 00
~ 01N .-~
N V' N 00OD
N M N ~ O1a'
M .--i.-~N O N
r .-rN ~ Tt'N
Q1r1r-i'-I.--Ir-i
O v~O~m
m n m cor m
r o~a~ .-~0 0
o r o N N ~r
a~o M u o rn
N M M M M M
O O O O O O
N N N N N N
-i.-a
Q'~ b~ ~ U~CT
..a
C C C C C C
O O O O O O
U U U U U U
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO 00122139 PCTNS99I23535
43
c roro b
o .c.c .r~
~
.v.G C C
y ~ T T
ro
,
a m ~n
i
..,
w ~ N N
.~ v b
,, .r,.r.,.'.,
c ~
~ ue x
' X '.
D S
~
.y ?~T 7r
.-I.-1
n
a a. ,
ro
,n
O
sa
rmn
-ao v~
N ~-1N
1
I
N
O 6 E 6
U ~ G C
fT t3~
C G C
N 4
i~ O O O
ro
~ r-1
U7~N N N
N C Q' cr
~ N N N
,Q
O O O
ro O~N
ro N O a'
N ~ttm ~1
~I7O~ N
N p 00O N
C N
~ ~ ~ N '-iN
r
l
171
~y b O
' ~~
N
Q) .-a,~ c
O
b
,1 ,--m U
a~ ~ w
,.. p n o ~ w
CI N a' O ~ c~
N N r-I
GI
al
N
W
N
e! N ~
!E
.-aN O
(n .-1
N ,O O O ,--1
'C1
y ~ I I
r oo U r m o0
~
ql ~ .-~.-,
to 1 .-r.-~r, v~tT O~
a
~ 1 ~ ~ .~ ~ "~"'1~.-i
01 . ~ N ~
.-
p, v a . C C C C
In C C
vl O O ~ U
U U
H b ~ U U .rl
SUBSTITUTE SHEET (RULE 26)
CA 02346499 2001-04-09
WO 00/22139 PCT/US99/23535
44
Table: 7. ORF data summary from A2 insert
c M a
m O cfl
r
o m i . N
.
o m
,p M ~ ~ M
~ ~ m
Y.. C ~
N '
. Q) gyp V ~ ~ ~ ~
,C O
~ O ~ ~ ~
~
0 c~
, ~ ~ ~ O
0
N tl O M T1 l~j
~ ~
~ o
Q ~ c c ~ vi
~ ~ a ~
N ~ . r c a. r r c~i
C i~ ~ O 00
'
'
_ ~ ~ f0 N O O
N ~ LA O
O M O (0 E Q (D
(O tn ~ N ~
0~7 ~ a N O ~ O ~ Oi
< N
' i
L N j, t Qi n3
t. n t,.
M L
' N O N O _ O ~! ~ ~ ~ O
~ ~ t
tn
4
t N N . t N
U - C n o n
~I'
'~t' m ~ .- o c c~ :o r.
m v ~ ~ ~
v~
- ~
C
~ V ~ C >. ~ C ~p ~ r
I~ ~ fa O pj ~
if. > ~ N t N O ~ ~ ~ O N
N f9 0 ~ ~
' N , ~ U t O ( ~ ~ C
:i N O /) ~ a
U
' O ~ ~ V ( N
O ~ ~ ~ 0
O ~ z '~ C_ p ~ .C LL' ici ~ Z O.
~' ~ p C -O CrD ~ ~j
N
'tf.X ~ ~X E .C N N ~ O O ~ ~ U
t0 >. V' O .C C tn
d O N (A Y 'O ~ M r ~ Z
t9 d N ..'..D ~ N
O N N
N ~0 N N N (0
C N U ~
N N t4
G7 ~C C ~ ~p O N O
tn p
_ p O ~ .N V C 'a rn N O OO O
O O f0 ~
. . O ~ ~ N O N p f0 f9
C ~ ~
flV ~ ~ ~ N ~ N . '~ _ 7
C U Q
O
. C . f0 77
C
O C.n z ~ O O ~ >' C ~ ~ OO
N ~
'~
~ ~ ' ' C C ~ Y a N a m NN N
O
Q.- ~ O t9 ~X . , cn
v .
y ~ E 0. O ~,
O. O
V f~ O
a
0 ~
0 h I'- ~ 00 cr M ~ Q
0 M
V .- r- r- T .- cO r~r-
~ r ~ ~ ~n
~'' f~ O O O N N N MO N
,
~. I ~ V N N ~" N
.
N V . ,
r
d. ~ tn ~ '~ tl~ O M OO lf7
V ~ tt'
+' s ~ ~' O O~ OD I~.t M .COtn
N ' tn 00
ct O N O N Oy I~ U OM CO
u7 N I~ r' CO O M O -'Oa~
N ' O W h
-
'
r M N ~f O c0 r- O -r-
t1 r-
a ~ U U U ~ ~c U c
~
c Q
V o r y w w . s w w ~ w
0 0 0 W 0 0 00 0 0 00 0
0 0
0
0 Wit' I~. f~ d' ~ Cfl O (DO M 1~NCO
O tn ~ o0
O O '~t O ~!' tt 00 1~- 00 (OCO~ tn'ctO (D
CO tn
O O 07 1~- 00 00 c0O O) 07OO O
O 00
'tf'~ 00 ~ ~- I~ OO ~ (Dt~I~
M Q7 N
M M ~.,~ ~7 ~. ~ CO NN ~ M ~~
O M
1~ h r- M CO CO OM O V'V'M N
f~ tt~
n T T V r- ~r- I~
N r'
N tt 07 N--N M i~~ O
of ~ M
CO tn
r 00 00 OO M ~ h~ M
~
M f cr r O CO V''~tO COON
M ~ ~
M c0 a0 ~ ~ N N M M MM M
M O~
M
CO ~ ~ O (fl V' M ~O N o0.-~ (p
O M
' CO O e- O tn h O tnN 00 N CO~-
O O Ln
fp (p r ~ (~ M O Nf~O i-CO(O
r M 00
O ~ O M tn O 00T T M M~ 1~
~
N NM M M MM M
N Wit' ~ !~ 00 O O~ N M c1tn
M (O r-r-~ e-~(O
SUBSTITUTE SHEET (RULE 26~